6.3 OML User Tasks for Downloading an Available Conda Environment
Once user ADMIN installs the environment in Object Storage in the Autonomous Database, as an OML user, you can download, activate, and use it in Python paragraphs in notebooks and with embedded execution.
List all environments persisted in Object Storage
Get the list of environments saved in Object Storage using the
list-saved-envs command.
%conda
list-saved-envs Get information on a named environment persisted in Object Storage
Provide the environment name as an argument to the -e
parameter and request information on the environment.
%conda
list-saved-envs -e sbenvThe output is similar to the following:
{
"name": "sbenv",
"size": "1.2 GiB",
"description": "Conda environment with seaborn",
"tags": {
"application": "OML4PY"
},
"number_of_installed_packages": 60
}Download and activate the environment
Use the download command to download an
environment from Object Storage. To activate the downloaded
environment, use the activate command.
Note:
The paragraph that contains the download command must be the first paragraph in the notebook.%conda
download sbenv
activate sbenvDownloading conda environment sbenv
Download successful for conda environment sbenvList the packages available in the environment
Get the list of all the packages in an active environment using the
list command.
%conda
listThe output is similar to the following:
# packages in environment at /u01/.conda/envs/sbenv:
#
# Name Version Build Channel
blas 1.0 mkl
bottleneck 1.3.5 py39h7deecbd_0
brotli 1.0.9 h5eee18b_7
brotli-bin 1.0.9 h5eee18b_7
ca-certificates 2022.07.19 h06a4308_0
certifi 2022.9.14 py39h06a4308_0
cycler 0.11.0 pyhd3eb1b0_0
dbus 1.13.18 hb2f20db_0
expat 2.4.4 h295c915_0
fftw 3.3.9 h27cfd23_1
fontconfig 2.13.1 h6c09931_0
fonttools 4.25.0 pyhd3eb1b0_0
freetype 2.11.0 h70c0345_0
giflib 5.2.1 h7b6447c_0
glib 2.69.1 h4ff587b_1
gst-plugins-base 1.14.0 h8213a91_2
gstreamer 1.14.0 h28cd5cc_2
icu 58.2 he6710b0_3
intel-openmp 2021.4.0 h06a4308_3561
jpeg 9e h7f8727e_0
kiwisolver 1.4.2 py39h295c915_0
lcms2 2.12 h3be6417_0
ld_impl_linux-64 2.38 h1181459_1
lerc 3.0 h295c915_0
libbrotlicommon 1.0.9 h5eee18b_7
libbrotlidec 1.0.9 h5eee18b_7
libbrotlienc 1.0.9 h5eee18b_7
libdeflate 1.8 h7f8727e_5
libffi 3.3 he6710b0_2
libgcc-ng 11.2.0 h1234567_1
libgfortran-ng 11.2.0 h00389a5_1
libgfortran5 11.2.0 h1234567_1
libpng 1.6.37 hbc83047_0
libstdcxx-ng 11.2.0 h1234567_1
libtiff 4.4.0 hecacb30_0
libuuid 1.0.3 h7f8727e_2
libwebp 1.2.2 h55f646e_0
libwebp-base 1.2.2 h7f8727e_0
libxcb 1.15 h7f8727e_0
libxml2 2.9.14 h74e7548_0
lz4-c 1.9.3 h295c915_1
matplotlib 3.5.2 py39h06a4308_0
matplotlib-base 3.5.2 py39hf590b9c_0
mkl 2021.4.0 h06a4308_640
mkl-service 2.4.0 py39h7f8727e_0
mkl_fft 1.3.1 py39hd3c417c_0
mkl_random 1.2.2 py39h51133e4_0
munkres 1.1.4 py_0
ncurses 6.3 h5eee18b_3
numexpr 2.8.3 py39h807cd23_0
numpy 1.22.3 py39he7a7128_0
numpy-base 1.22.3 py39hf524024_0
openssl 1.1.1q h7f8727e_0
packaging 21.3 pyhd3eb1b0_0
pandas 1.4.4 py39h6a678d5_0
pcre 8.45 h295c915_0
pillow 9.2.0 py39hace64e9_1
pip 22.1.2 py39h06a4308_0
pyparsing 3.0.9 py39h06a4308_0
pyqt 5.9.2 py39h2531618_6
python 3.9.0 hdb3f193_2
python-dateutil 2.8.2 pyhd3eb1b0_0
pytz 2022.1 py39h06a4308_0
qt 5.9.7 h5867ecd_1
readline 8.1.2 h7f8727e_1
scipy 1.7.3 py39h6c91a56_2
seaborn 0.11.2 pyhd3eb1b0_0
setuptools 63.4.1 py39h06a4308_0
sip 4.19.13 py39h295c915_0
six 1.16.0 pyhd3eb1b0_1
sqlite 3.39.2 h5082296_0
tk 8.6.12 h1ccaba5_0
tornado 6.2 py39h5eee18b_0
tzdata 2022c h04d1e81_0
wheel 0.37.1 pyhd3eb1b0_0
xz 5.2.5 h7f8727e_1
zlib 1.2.12 h5eee18b_3
zstd 1.5.2 ha4553b6_0Example 6-1 Create a visualization using seaborn
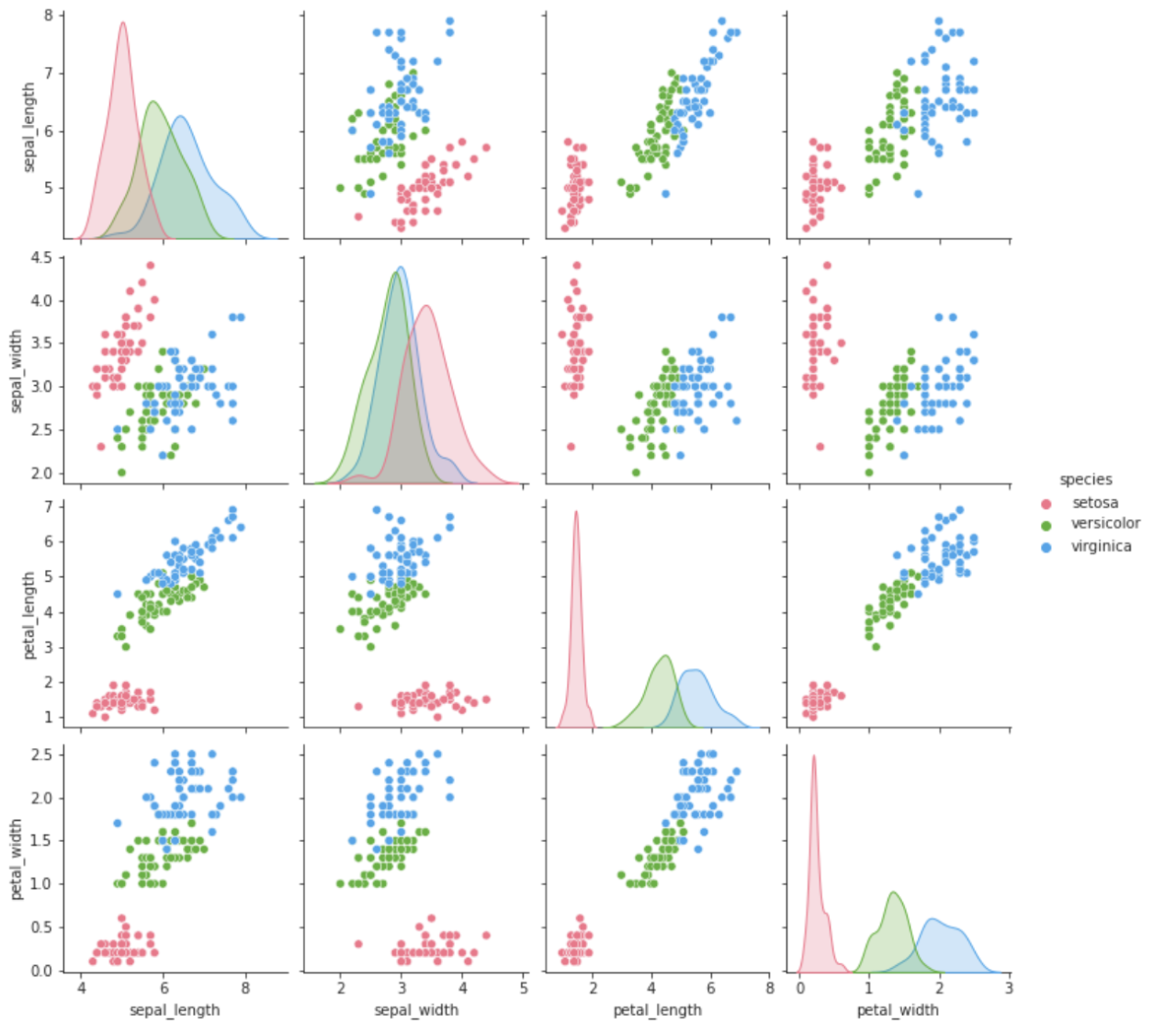
The following example shows the use of the available
packages in the installed and activated environment. It imports
pandas, seaborn, and matplotlib packages
and loads the iris dataset from the seaborn library
as a pandas dataframe. The pairplot seaborn
function plots the pair-wise relationship between all the variables
of the dataset.
%python
import pandas as pd
import seaborn as sb
from matplotlib import pyplot as plt
df = sb.load_dataset('iris')
sb.set_style("ticks")
sb.pairplot(df,hue = 'species',diag_kind = "kde",kind = "scatter",palette = "husl")
plt.show()The output of the example is the following.
Example 6-2 Create a string representation of the function and save it to the OML4Py script repository
With OML4Py, functions are saved to the script repository
using their string definition representation so they can be run in
embedded Python execution. Create a function
sb_plot, after verifying the function
behaves as expected, provide it as a string (within triple quotes
for formatting), and save it to the OML4Py script repository. Use
the oml.script.create function to store a single
user-defined Python function in the OML4Py script repository. The
parameter "sb_plot" is a string that specifies the
name of the user-defined function. The parameter
func=sb_plot is the Python function to
run.
%python
sb_plot = """def sb_plot():
import pandas as pd
import seaborn as sb
from matplotlib import pyplot as plt
df = sb.load_dataset('iris')
sb.set_style("ticks")
sb.pairplot(df,hue = 'species',diag_kind = "kde",kind = "scatter",palette = "husl")
plt.show()"""
oml.script.create("sb_plot", func=sb_plot)Use the Python API for embedded Python execution to run the user-defined Python function you saved in the script repository.
%python
oml.do_eval(func="sb_plot", graphics=True)The output of the example is the following:
Deactivate the current environment
Use the deactivate command to deactivate an
enviroment.
Note:
At a given time, only one active environment is supported. So, a newly activated environment would replace an old environment. As a best practice, deactivate an environment before logging off.%conda
deactivateConda environment deactivatedParent topic: Install Third-Party Packages