6 Using Oracle R Enterprise Embedded R Execution
Embedded R execution in Oracle R Enterprise enables you to invoke R scripts in R sessions that run on the Oracle Database server. This chapter discusses embedded R execution in the following topics:
About Oracle R Enterprise Embedded R Execution
In Oracle R Enterprise, embedded R execution is the ability to store R scripts in the Oracle Database R script repository and to invoke such scripts. When invoked, a script executes in one or more R engines that run on the database server and that are dynamically started and managed by the database. Oracle R Enterprise provides both an R interface and a SQL interface for embedded R execution. From the same R script you can get structured data, an XML representation of R objects and images, and even PNG images through a BLOB column in a database table.
This section has the following topics:
Benefits of Embedded R Execution
Embedded R execution has the following benefits:
-
Eliminates moving data from the Oracle Database server to your local R session.
As well as being more secure, the transfer of database data between Oracle Database and an internal R engine is much faster than to a separate client R engine.
-
Uses the database server to start, manage, and control the execution of R scripts in R engines running on the server.
-
Leverages the memory and processing power of the database server machine for R engine execution, which provides better scalability and performance.
-
Enables data-parallel and task-parallel execution of user-defined R functions that correspond to special cases of Hadoop Map-Reduce jobs.
-
Provides parallel simulations capability.
-
Allows the use of open source CRAN packages in R scripts running on the database server.
-
Provides the ability to develop and operationalize comprehensive scripts for analytical applications in a single step, without leaving the R environment.
You can directly integrate R scripts used in exploratory analysis into application tasks. You can also immediately invoke R scripts in production to drastically reduce time to market by eliminating porting and enabling instantaneous updates of changes to application code.
-
Executing R scripts from SQL enables integration of R script results with Oracle Business Intelligence Enterprise Edition (OBIEE), Oracle BI Publisher, and other SQL-enabled tools for structured data, R objects, and images.
APIs for Embedded R Execution
Oracle R Enterprise provides R and SQL application programming interfaces for embedded R execution. Table 6-1 provides a summary of the embedded R execution functions and the R script repository functions available. The function f refers to the user-defined R closure, or script, that is provided as either an R function object or a named R function in the database R script repository.
Table 6-1 R and SQL APIs for Embedded R Execution
| R API | SQL API | Description |
|---|---|---|
|
|
|
Executes f with no automatic transfer of data. |
|
|
|
Executes f by passing all rows of the provided input |
|
|
rqGroupEval This function must be explicitly defined by the user. |
Executes f by partitioning data according to the values of a grouping column. Provides each data partition as a |
|
|
|
Executes f by passing a specified number of rows (a chunk) of the provided input |
|
|
No equivalent. |
Executes f with no automatic transfer of data but provides the index of the invocation, 1 through n, where n is the number of times to invoke the function. Supports parallel execution of each f invocation in the pool of R engines running on the database server. |
|
|
|
Loads the provided R function into the R script repository with the provided name. |
|
|
|
Removes the named R function from the R script repository. |
Security Considerations for Scripts
Because R scripts allow access to the database server, the creation of scripts must be controlled. The RQADMIN role is a collection of Oracle Database privileges that a user must have to create scripts and store them in the Oracle Database R script repository or drop scripts from the repository.
The installation of Oracle R Enterprise on the database server creates the RQADMIN role. The role must be explicitly granted to a user. To grant RQADMIN to a user, start SQL*Plus as sysdba and enter a GRANT statement such as the following, which grants the role to the user RQUSER:
GRANT RQADMIN to RQUSER
Note:
You should grant RQADMIN only to those users who need it.Support for Parallel Execution
Some of the Oracle R Enterprise embedded R execution functions support the use of parallel execution in the database. The ore.groupApply, ore.rowApply, rq.groupEval, and rq.rowEval functions support data-parallel execution and the ore.indexApply function supports task-parallel execution. This parallel execution capability enables a script to take advantage of high-performance computing hardware such as an Oracle Exadata Database Machine.
The parallel argument of the ore.groupApply, ore.rowApply, and ore.indexApply functions specifies the degree of parallelism to use in the embedded R execution. The value of the argument can be one of the following:
-
A positive integer greater than or equal to
2for a specific degree of parallelism -
FALSEor1for no parallelism -
TRUEfor the default parallelism of thedataargument -
NULLfor the database default for the operation
The default value of the argument is the value of the global option ore.parallel or FALSE if ore.parallel is not set.
A user-defined R function invoked using ore.doEval or ore.tableApply is not executed in parallel. The function executes in a single R engine.
For the rq.groupEval, and rq.rowEval functions, the degree of parallelism is specified by a PARALLEL hint in the input cursor argument.
In data-parallel execution for the ore.groupApply and rq.groupEval functions, one or more R engines perform the same R function, or task, on different partitions of data. This functionality enables the building of large numbers of models, for example building tens or hundreds of thousands of predictive models, one model per customer.
In data-parallel execution for the ore.rowApply and rq.rowEval functions, one or more R engines perform the same R function on disjoint chunks of data. This functionality enables scalable model scoring and predictions on large data sets.
In task-parallel execution for the ore.indexApply function, one or more R engines perform the same or different calculations, or task. A number, associated with the index of the execution, is provided to the function. This functionality is valuable in a variety of operations, such as in performing simulations.
Oracle Database handles the management and control of potentially multiple R engines at the database server, automatically partitioning and passing data to R engines executing in parallel. It ensures that all of the R function executions for all of the partitions complete; if not, the Oracle R Enterprise function returns an error. The result from the execution of each user-defined embedded R function is gathered in an ore.list. This list remains in the database until the user requires the result.
Embedded R execution also allows for data-parallel execution of user-defined R functions that may use functions from an open source R package from The Comprehensive R Archive Network (CRAN) or other third-party R package. However, third-party packages do not leverage in-database parallelism and are subject to the parallelism constraints of R. Third-party packages can benefit from the data-parallel and task-parallel execution supported in embedded R execution.
See Also:
"Oracle R Enterprise Global Options"Installing a Third-Party Package for Use in Embedded R Execution
Embedded R execution allows the use of CRAN or other third-party packages in user-defined R functions executed on the Oracle Database server. To use a third-party package in embedded R execution, the package must be installed on the database server. If you are going to use the package from the R interface for embedded R execution, then the package must also be installed on the client, as well. To avoid incompatibilities, you must install the same version of the package on both the client and server machines.
An Oracle Database Administrator (DBA) can install a package on a database server so that it can be used by embedded R execution functions or by any R user. The DBA can install a package on a single database server or on multiple database servers.
A DBA would typically do the following:
-
Download and install the package from CRAN. Downloading a package from CRAN requires an Internet connection.
-
In an Oracle R Enterprise session running on the server, load the package. Verify that the package is installed correctly by using a function in the package.
To install a package on a single database server, do one of the following:
-
In an Oracle R Enterprise session running on the server, invoke the
install.packagesfunction, as shown in Example 6-1. The function downloads the package and installs dependencies automatically. -
Download the package source from CRAN using
wget. If the package depends on any packages that are not in the R distribution in use, then download those packages, also.From the operating system command line, use the
ORE CMD INSTALLcommand to install the package or packages in the same location as the Oracle R Enterprise packages, which is$ORACLE_HOME/R/library. See Example 6-2.
To install a package, and any dependent packages, on multiple database servers, such as those in an Oracle Real Application Clusters (Oracle RAC) or a multinode Oracle Exadata Database Machine environment, use the Exadata Distributed Command Line Interface (DCLI) utility, as shown in Example 6-3. For detailed instructions on using DCLI to install packages, see Oracle R Enterprise Installation and Administration Guide.
To verify that the package is installed correctly, load the package and use a function in the package, as shown in Example 6-4.
Example 6-1 invokes the install.packages function to download the C50 package from CRAN and to install it. The C50 package contains functions for creating C5.0 decision trees and rule-based models for pattern recognition.
Example 6-1 Installing a Package for a Single Database in an Oracle R Enterprise Session
install.packages("c50")
The output Example 6-1 is almost identical to the output of the ORE CMD INSTALL command in Example 6-2.
Example 6-2 demonstrates downloading the C50 package from CRAN and installing it with ORE CMD INSTALL from a Linux command line.
Example 6-2 Installing a Package for a Single Database from the Command Line
wget http://cran.r-project.org/src/contrib/C50_0.1.0-19.tar.gz ORE CMD INSTALL C50_0.1.0-19.tar.gz
Listing for Example 6-2
$ wget http://cran.r-project.org/src/contrib/C50_0.1.0-19.tar.gz
# The output of wget is not shown.
$ ORE CMD INSTALL C50_0.1.0-19.tar.gz
* installing to library '/example/dbhome_1/R/library'
* installing *source* package 'C50' ...
** package 'C50' successfully unpacked and MD5 sums checked
checking for gcc... gcc
checking whether the C compiler works... yes
checking for C compiler default output file name... a.out
checking for suffix of executables...
checking whether we are cross compiling... no
checking for suffix of object files... o
checking whether we are using the GNU C compiler... yes
checking whether gcc accepts -g... yes
checking for gcc option to accept ISO C89... none needed
configure: creating ./config.status
config.status: creating src/Makevars
** libs
gcc -m64 -std=gnu99 -I/usr/include/R -DNDEBUG -DNDEBUG -I/usr/local/include -ffloat-store -g -fpic -g -O2 -c attwinnow.c -o attwinnow.o
gcc -m64 -std=gnu99 -I/usr/include/R -DNDEBUG -DNDEBUG -I/usr/local/include -ffloat-store -g -fpic -g -O2 -c classify.c -o classify.o
gcc -m64 -std=gnu99 -I/usr/include/R -DNDEBUG -DNDEBUG -I/usr/local/include -ffloat-store -g -fpic -g -O2 -c confmat.c -o confmat.o
gcc -m64 -std=gnu99 -I/usr/include/R -DNDEBUG -DNDEBUG -I/usr/local/include -ffloat-store -g -fpic -g -O2 -c construct.c -o construct.o
gcc -m64 -std=gnu99 -I/usr/include/R -DNDEBUG -DNDEBUG -I/usr/local/include -ffloat-store -g -fpic -g -O2 -c contin.c -o contin.o
gcc -m64 -std=gnu99 -I/usr/include/R -DNDEBUG -DNDEBUG -I/usr/local/include -ffloat-store -g -fpic -g -O2 -c discr.c -o discr.o
gcc -m64 -std=gnu99 -I/usr/include/R -DNDEBUG -DNDEBUG -I/usr/local/include -ffloat-store -g -fpic -g -O2 -c formrules.c -o formrules.o
gcc -m64 -std=gnu99 -I/usr/include/R -DNDEBUG -DNDEBUG -I/usr/local/include -ffloat-store -g -fpic -g -O2 -c formtree.c -o formtree.o
gcc -m64 -std=gnu99 -I/usr/include/R -DNDEBUG -DNDEBUG -I/usr/local/include -ffloat-store -g -fpic -g -O2 -c getdata.c -o getdata.o
gcc -m64 -std=gnu99 -I/usr/include/R -DNDEBUG -DNDEBUG -I/usr/local/include -ffloat-store -g -fpic -g -O2 -c getnames.c -o getnames.o
gcc -m64 -std=gnu99 -I/usr/include/R -DNDEBUG -DNDEBUG -I/usr/local/include -ffloat-store -g -fpic -g -O2 -c global.c -o global.o
gcc -m64 -std=gnu99 -I/usr/include/R -DNDEBUG -DNDEBUG -I/usr/local/include -ffloat-store -g -fpic -g -O2 -c hash.c -o hash.o
gcc -m64 -std=gnu99 -I/usr/include/R -DNDEBUG -DNDEBUG -I/usr/local/include -ffloat-store -g -fpic -g -O2 -c hooks.c -o hooks.o
gcc -m64 -std=gnu99 -I/usr/include/R -DNDEBUG -DNDEBUG -I/usr/local/include -ffloat-store -g -fpic -g -O2 -c implicitatt.c -o implicitatt.o
gcc -m64 -std=gnu99 -I/usr/include/R -DNDEBUG -DNDEBUG -I/usr/local/include -ffloat-store -g -fpic -g -O2 -c info.c -o info.o
gcc -m64 -std=gnu99 -I/usr/include/R -DNDEBUG -DNDEBUG -I/usr/local/include -ffloat-store -g -fpic -g -O2 -c mcost.c -o mcost.o
gcc -m64 -std=gnu99 -I/usr/include/R -DNDEBUG -DNDEBUG -I/usr/local/include -ffloat-store -g -fpic -g -O2 -c modelfiles.c -o modelfiles.o
gcc -m64 -std=gnu99 -I/usr/include/R -DNDEBUG -DNDEBUG -I/usr/local/include -ffloat-store -g -fpic -g -O2 -c p-thresh.c -o p-thresh.o
gcc -m64 -std=gnu99 -I/usr/include/R -DNDEBUG -DNDEBUG -I/usr/local/include -ffloat-store -g -fpic -g -O2 -c prune.c -o prune.o
gcc -m64 -std=gnu99 -I/usr/include/R -DNDEBUG -DNDEBUG -I/usr/local/include -ffloat-store -g -fpic -g -O2 -c rc50.c -o rc50.o
gcc -m64 -std=gnu99 -I/usr/include/R -DNDEBUG -DNDEBUG -I/usr/local/include -ffloat-store -g -fpic -g -O2 -c redefine.c -o redefine.o
gcc -m64 -std=gnu99 -I/usr/include/R -DNDEBUG -DNDEBUG -I/usr/local/include -ffloat-store -g -fpic -g -O2 -c rsample.c -o rsample.o
gcc -m64 -std=gnu99 -I/usr/include/R -DNDEBUG -DNDEBUG -I/usr/local/include -ffloat-store -g -fpic -g -O2 -c rulebasedmodels.c -o rulebasedmodels.o
gcc -m64 -std=gnu99 -I/usr/include/R -DNDEBUG -DNDEBUG -I/usr/local/include -ffloat-store -g -fpic -g -O2 -c rules.c -o rules.o
gcc -m64 -std=gnu99 -I/usr/include/R -DNDEBUG -DNDEBUG -I/usr/local/include -ffloat-store -g -fpic -g -O2 -c ruletree.c -o ruletree.o
gcc -m64 -std=gnu99 -I/usr/include/R -DNDEBUG -DNDEBUG -I/usr/local/include -ffloat-store -g -fpic -g -O2 -c siftrules.c -o siftrules.o
gcc -m64 -std=gnu99 -I/usr/include/R -DNDEBUG -DNDEBUG -I/usr/local/include -ffloat-store -g -fpic -g -O2 -c sort.c -o sort.o
gcc -m64 -std=gnu99 -I/usr/include/R -DNDEBUG -DNDEBUG -I/usr/local/include -ffloat-store -g -fpic -g -O2 -c strbuf.c -o strbuf.o
gcc -m64 -std=gnu99 -I/usr/include/R -DNDEBUG -DNDEBUG -I/usr/local/include -ffloat-store -g -fpic -g -O2 -c subset.c -o subset.o
gcc -m64 -std=gnu99 -I/usr/include/R -DNDEBUG -DNDEBUG -I/usr/local/include -ffloat-store -g -fpic -g -O2 -c top.c -o top.o
gcc -m64 -std=gnu99 -I/usr/include/R -DNDEBUG -DNDEBUG -I/usr/local/include -ffloat-store -g -fpic -g -O2 -c trees.c -o trees.o
gcc -m64 -std=gnu99 -I/usr/include/R -DNDEBUG -DNDEBUG -I/usr/local/include -ffloat-store -g -fpic -g -O2 -c
update.c -o update.o
gcc -m64 -std=gnu99 -I/usr/include/R -DNDEBUG -DNDEBUG -I/usr/local/include -ffloat-store -g -fpic -g -O2 -c utility.c -o utility.o
gcc -m64 -std=gnu99 -I/usr/include/R -DNDEBUG -DNDEBUG -I/usr/local/include -ffloat-store -g -fpic -g -O2 -c xval.c -o xval.o
gcc -m64 -std=gnu99 -shared -L/usr/local/lib64 -o C50.so attwinnow.o classify.o confmat.o construct.o contin.o discr.o formrules.o formtree.o getdata.o getnames.o global.o hash.o hooks.o implicitatt.o info.o mcost.o modelfiles.o p-thresh.o prune.o rc50.o redefine.o rsample.o rulebasedmodels.o rules.o ruletree.o siftrules.o sort.o strbuf.o subset.o top.o trees.o update.o utility.o xval.o -L/usr/lib64/R/lib -lR
installing to /example/dbhome_1/R/library/C50/libs
** R
** data
** preparing package for lazy loading
** help
*** installing help indices
converting help for package 'C50'
finding HTML links ... done
C5.0 html
C5.0Control html
churn html
predict.C5.0 html
summary.C5.0 html
varImp.C5.0 html
** building package indices
** testing if installed package can be loaded
* DONE (C50)
Example 6-3 shows the DLCI command for installing the C50 package.
Example 6-3 Installing a Package Using DCLI
dcli -g nodes -l oracle R CMD INSTALL C50_0.1.0-19.tar.gz
The dcli -g flag designates a file containing a list of nodes to install on, and the -l flag specifies the user ID to use when executing the commands. For more information on using DCLI, see Oracle R Enterprise Installation and Administration Guide.
Example 6-4 shows starting R, connecting to Oracle R Enterprise on the server, loading the C50 package, and using a function in the package. The example starts R by executing the ORE command from the Linux command line. The example connects to Oracle R Enterprise and then loads the C50 package. It invokes the demo function to look for example programs in the package. Because the package does not have examples, Example 6-4 then gets help for the C5.0 function. The example invokes example code from the help.
Example 6-4 Using a C50 Package Function
ORE
library(ORE)
ore.connect(user = "RQUSER", sid = "orcl", host = "myhost",
password = "rquserStrongPassword", port = 1521, all=TRUE)
library(C50)
demo(package = "C50")
?C5.0
data(churn)
treeModel <- C5.0(x = churnTrain[, -20], y = churnTrain$churn)
treeModel
Listing for Example 6-4
$ ORE
R> library(ORE)
Loading required package: OREbase
Attaching package: 'OREbase'
The following objects are masked from 'package:base':
cbind, data.frame, eval, interaction, order, paste, pmax, pmin,
rbind, table
Loading required package: OREembed
Loading required package: OREstats
Loading required package: MASS
Loading required package: OREgraphics
Loading required package: OREeda
Loading required package: OREmodels
Loading required package: OREdm
Loading required package: lattice
Loading required package: OREpredict
Loading required package: ORExml
> ore.connect(user = "RQUSER", sid = "orcl", host = "myhost",
+ password = "rquserStrongPassword", port = 1521, all=TRUE)
Loading required package: ROracle
Loading required package: DBI
R> library(C50)
R> demo(package = "C50")
no demos found
R> ?C5.0 # Output not shown.
R> data(churn)
R> treeModel <- C5.0(x = churnTrain[, -20], y = churnTrain$churn)
R> treeModel
Call:
C5.0.default(x = churnTrain[, -20], y = churnTrain$churn)
Classification Tree
Number of samples: 3333
Number of predictors: 19
Tree size: 27
Non-standard options: attempt to group attributes
See Also:
R Interface for Embedded R Execution
Oracle R Enterprise provides functions that invoke R scripts that run in one or more R engines that are embedded in the Oracle database. Other functions create and store an R script in a database R script repository or drop a script from the repository. This section describes these functions in the following topics:
Arguments for Functions that Run Scripts
The Oracle R Enterprise embedded R execution functions ore.doEval, ore.tableApply, ore.groupApply, ore.rowApply, and ore.indexApply have arguments that are common to some or all of the functions. Some of the functions also have an argument that is unique to the function.
This section describes the arguments in the following topics:
See Also:
-
For function signatures and more details about function arguments, see the online help displayed by invoking
help(ore.doEval) -
For examples of the use of the arguments, see "Using the ore.doEval Function" and the other topics on using the embedded R execution functions
Input Function to Execute
The embedded R execution functions all require a function to apply during the execution of the script. You specify the input function with one of the following mutually exclusive arguments:
-
FUN -
FUN.NAME
The FUN argument takes a function object as a directly specified function or as one assigned to an R variable. Only a user with the RQADMIN role can use the FUN argument when invoking an embedded R function.
The FUN.NAME argument specifies a script that is stored in the R script repository. A stored script contains the function to apply when the script runs. Any Oracle R Enterprise user can use the FUN.NAME argument when invoking an embedded R function.
Note:
The advanced Oracle R Enterprise analytics functions in theOREmodels package, ore.glm, ore.lm, and ore.neural, use the embedded R execution framework internally and cannot be used in embedded R execution functions.Optional and Control Arguments
All of the embedded R execution functions take optional arguments, which can be named or not. Oracle R Enterprise passes user-defined optional arguments to the input function. You can pass any number of optional arguments to the input function, including complex R objects such as models.
Arguments that start with ore. are special control arguments. Oracle R Enterprise does not pass them to the input function, but instead uses them to control what happens before or after the execution of that function. The following control arguments are supported:
-
ore.connectcontrols whether to automatically connect to Oracle R Enterprise inside the embedded R execution function. This is equivalent to doing anore.connectcall with the same credentials as the client session. The default value isFALSE.If an automatic connection is enabled, the following functionality occurs:
-
The embedded R script is connected to the database.
-
The connection has the same credentials as the session that invokes the embedded R SQL function.
-
The script runs in an autonomous transaction.
-
ROracle queries can work with the automatic connection.
-
Oracle R Enterprise transparency layer functionality is enabled in the embedded script.
-
-
ore.dropcontrols the input data. If the option value isTRUE, a one columndata.frameis converted to avector. The default value isTRUE. -
ore.envAsEmptyenvcontrols whether an environment referenced in an object is replaced with an empty environment during serialization. Some types of input parameters and returned objects, such aslistandformula, are serialized before being saved to the database. If the control argument value isTRUE, then the referenced environment in the object is replaced with an empty environment whose parent is.GlobalEnvand the objects in the original referenced environment are not serialized. In some cases, this can significantly reduce the size of serialized objects. If the control argument value isFALSE, then all of the objects in the referenced environment are serialized and can be unserialized and recovered later. The default value is regulated by the global optionore.envAsEmptyenv. -
ore.na.omitcontrols the handling of missing values in the input data. If you specifyore.na.omit = TRUE, then rows or vector elements, depending on theore.dropsetting, that contain missing values are removed from the input data. If all of the rows in a chunk contain missing values, then the input data for that chunk will be an emptydata.frameorvector. The default value isFALSE. -
ore.graphicscontrols whether to start a graphical driver and look for images. The default value isTRUE. -
ore.png.*specifies additional arguments for thepnggraphics driver ifore.graphicsisTRUE. The naming convention for these arguments is to add anore.png.prefix to the arguments of thepngfunction. For example, ifore.png.heightis supplied, argumentheightis passed to thepngfunction. If not set, the standard default values for thepngfunction are used.
See Also:
For more details about control arguments, see the online help displayed by invokinghelp(ore.doEval)Structure of Return Value
Another argument that applies to all of the embedded R execution functions is FUN.VALUE. If the FUN.VALUE argument is NULL, then the ore.doEval and ore.tableApply function can return a serialized R object as an ore.object class object, and the ore.groupApply, ore.indexApply, and ore.rowApply functions return an ore.list object. However, if you specify a data.frame or an ore.frame with the FUN.VALUE argument, then the function returns an ore.frame that has the structure of the specified data.frame or ore.frame object.
Input Data
The ore.doEval and ore.indexApply functions do not automatically receive any data from the database. They simply execute the function specified by the FUN or FUN.NAME argument. Any data needed by the input function is either generated within that function or explicitly retrieved from a data source such as Oracle Database, other databases, or flat files. The input function can load data from a file or a table using the ore.pull function or other transparency layer function.
The ore.tableApply, ore.groupApply, and ore.rowApply functions require a database table as input data. The table is represented by an ore.frame. You supply that data with an ore.frame object that you specify with the X argument, which is the first argument to the embedded R execution function. The embedded R execution function passes the ore.frame object to the user-defined input function as the first argument to that function.
Note:
The data represented by theore.frame object passed to the user-defined R function is copied from Oracle Database to the database server R engine. The R memory limitations apply. If your database server machine has 32 GB RAM and your data table is 64 GB, then Oracle R Enterprise cannot load the data into the R engine memory.Parallel Execution
The ore.groupApply, ore.indexApply, and ore.rowApply functions take the parallel argument. That argument specifies the degree of parallelism to use in the embedded R execution of the input function. See "Support for Parallel Execution".
Unique Arguments
The ore.groupApply, ore.indexApply, and ore.rowApply functions each take an argument unique to the function.
The ore.groupApply function takes the INDEX argument, which specifies the name of a column by which the rows of the input data are partitioned for processing by the input function.
The ore.indexApply function takes the times argument, which specifies the number of times to execute the input function.
The ore.rowApply function tales the rows argument, which specifies the number of rows to pass to each invocation of the input function.
Managing Scripts Using the R API
As mentioned in "Input Function to Execute," the embedded R execution functions can take a FUN.VALUE argument. That argument specifies the name of a script in the R script repository. To add a script to the repository, you invoke the ore.scriptCreate function. Scripts in the R script repository are also available through the SQL API for embedded R execution.
The ore.scriptDrop function removes the specified script from the R script repository.
Note:
Invoking theore.scriptCreate or ore.scriptDrop function requires the RQADMIN role. For more information, see "Security Considerations for Scripts".Both the ore.scriptCreate and ore.scriptDrop functions return an invisible NULL value if they succeed. If the function does not succeed in creating or dropping the script, it returns an error.
Example 6-5 first invokes ore.scriptDrop to ensure that the R script repository does not contain a script with the specified name. The example then invokes the ore.scriptCreate function to create the user-defined function named myRandomRedDots. The user-defined function accepts an argument, and it returns a data.frame object that has two columns and that plots 100 random normal values. The ore.scriptCreate invocation stores myRandomRedDots in the R script repository.
Example 6-5 Using the ore.scriptCreate and ore.scriptDrop Functions
ore.scriptDrop("myRandomRedDots")
ore.scriptCreate("myRandomRedDots", function(divisor = 100){
id <- 1:10
plot(1:100, rnorm(100), pch = 21, bg = "red", cex = 2 )
data.frame(id = id, val = id / divisor)
})
See Also:
-
"Using the ore.doEval Function" for examples that uses the function defined in Example 6-5
-
Example 6-13 for another example of using
ore.scriptCreateandore.scriptDrop
Using the ore.doEval Function
The ore.doEval function executes the specified input function using data that is generated by the input function. It returns an ore.frame object or a serialized R object as an ore.object object.
The syntax of the ore.doEval function is the following:
ore.doEval(FUN, ..., FUN.VALUE = NULL, FUN.NAME = NULL)
See Also:
"Arguments for Functions that Run Scripts" for descriptions of the arguments to functionore.doEvalIn Example 6-6, RandomRedDots gets a function that has an argument and that returns a data.frame object that has two columns and that plots 100 random normal values. The example then invokes ore.doEval function and passes it the RandomRedDots function object. The image is displayed at the client, but it is generated by the database server R engine that executed the RandomRedDots function.
Example 6-6 Using the ore.doEval Function
RandomRedDots <- function(divisor = 100){
id<- 1:10
plot(1:100, rnorm(100), pch = 21, bg = "red", cex = 2 )
data.frame(id=id, val=id / divisor)
}
ore.doEval(RandomRedDots)
Listing for Example 6-6
R> RandomRedDots <- function(divisor = 100){
+ id<- 1:10
+ plot(1:100, rnorm(100), pch = 21, bg = "red", cex = 2 )
+ data.frame(id=id, val=id / divisor)
+ }
R> ore.doEval(RandomRedDots)
id val
1 1 0.01
2 2 0.02
3 3 0.03
4 4 0.04
5 5 0.05
6 6 0.06
7 7 0.07
8 8 0.08
9 9 0.09
10 10 0.10
You can provide arguments to the input function as optional arguments to the doEval function. Example 6-7 invokes the doEval function with an optional argument that overrides the divisor argument of the RandomRedDots function.
Example 6-7 Using the ore.doEval Function with an Optional Argument
ore.doEval(RandomRedDots, divisor = 50)
Listing for Example 6-7
R> ore.doEval(RandomRedDots, divisor = 50) id val 1 1 0.02 2 2 0.04 3 3 0.06 4 4 0.08 5 5 0.10 6 6 0.12 7 7 0.14 8 8 0.16 9 9 0.18 10 10 0.20 # The graph displayed by the plot function is not shown.
If the input function is stored in the R script repository, then you can invoke the ore.doEval function with the FUN.NAME argument. Example 6-8 first invokes ore.scriptDrop to ensure that the R script repository does not contain a script with the name myRandomRedDots. The example adds the RandomRedDots function from Example 6-6 to the repository under the name myRandomRedDots. Example 6-8 invokes the ore.doEval function and specifies myRandomRedDots. The result is assigned to the variable res.
The return value of the RandomRedDots function is a data.frame but in Example 6-8 the ore.doEval function returns an ore.object object. To get back the data.frame object, the example invokes ore.pull to pull the result to the client R session.
Example 6-8 Using the ore.doEval Function with the FUN.NAME Argument
ore.scriptDrop("myRandomRedDots")
ore.scriptCreate("myRandomRedDots", RandomRedDots)
res <- ore.doEval(FUN.NAME = "myRandomRedDots", divisor = 50)
class(res)
res.local <- ore.pull(res)
class(res.local)
Listing for Example 6-8
R> ore.scriptDrop("myRandomRedDots")
R> ore.scriptCreate("myRandomRedDots", RandomRedDots)
R> res <- ore.doEval(FUN.NAME = "myRandomRedDots", divisor = 50)
R> class(res)
[1] "ore.object"
attr(,"package")
[1] "OREembed"
R> res.local <- ore.pull(res)
R> class(res.local)
[1] "data.frame"
To have the doEval function return an ore.frame object instead of an ore.object, use the argument FUN.VALUE to specify the structure of the result, as shown in Example 6-9.
Example 6-9 Using the ore.doEval Function with the FUN.VALUE Argument
res.of <- ore.doEval(FUN.NAME="myRandomRedDots", divisor = 50,
FUN.VALUE= data.frame(id = 1, val = 1))
class(res.of)
Listing for Example 6-9
R> res.of <- ore.doEval(FUN.NAME="myRandomRedDots", divisor = 50, + FUN.VALUE= data.frame(id = 1, val = 1)) R> class(res.of) [1] "ore.frame" attr(,"package") [1] "OREbase"
Example 6-10 demonstrates using the special optional argument ore.connect to connect to the database in the embedded R function, which enables the use of objects stored in a datastore. The example creates the RandomRedDots2 function object, which is the same as the RandomRedDots function from Example 6-6 except the RandomRedDots2 function has an argument that takes the name of a datastore. The example creates the myVar variable and saves it in the datastore named datastore_1. The example then invokes the doEval function and passes it the name of the datastore and passes the ore.connect control argument set to TRUE.
Example 6-10 Using the doEval Function with the ore.connect Argument
RandomRedDots2 <- function(divisor = 100, datastore.name = "myDatastore"){
id <- 1:10
plot(1:100, rnorm(100), pch = 21, bg = "red", cex = 2 )
ore.load(datastore.name) # Contains the numeric variable myVar.
data.frame(id = id, val = id / divisor, num = myVar)
}
myVar <- 5
ore.save(myVar, name = "datastore_1")
ore.doEval(RandomRedDots2, datastore.name = "datastore_1", ore.connect = TRUE)
Listing for Example 6-10
R> RandomRedDots2 <- function(divisor = 100, datastore.name = "myDatastore"){
+ id <- 1:10
+ plot(1:100, rnorm(100), pch = 21, bg = "red", cex = 2 )
+ ore.load(datastore.name) # Contains the numeric variable myVar.
+ data.frame(id = id, val = id / divisor, num = myVar)
+ }
R> ore.doEval(RandomRedDots2, datastore.name = "datastore_1", ore.connect = TRUE)
id val num
1 1 0.01 5
2 2 0.02 5
3 3 0.03 5
4 4 0.04 5
5 5 0.05 5
6 6 0.06 5
7 7 0.07 5
8 8 0.08 5
9 9 0.09 5
10 10 0.10 5
# The graph displayed by the plot function is not shown.
Using the ore.tableApply Function
The ore.tableApply function invokes an R script with an ore.frame as the input data. The ore.tableApply function passes the ore.frame to the user-defined input function as the first argument to that function. The ore.tableApply function returns an ore.frame object or a serialized R object as an ore.object object.
The syntax of the ore.tableApply function is the following:
ore.tableApply(X, FUN, ..., FUN.VALUE = NULL, FUN.NAME = NULL)
See Also:
-
"Arguments for Functions that Run Scripts" for descriptions of the arguments to function
ore.tableApply -
"Installing a Third-Party Package for Use in Embedded R Execution"
Example 6-11 uses the ore.tableApply function to build a Naive Bayes model on the iris data set. The naiveBayes function is in the e1071 package, which must be installed on both the client and database server machine R engines. As the first argument to the ore.tableApply function, the ore.push(iris) invocation creates a temporary database table and an ore.frame that is a proxy for the table. The second argument is the input function, which has as an argument dat. The ore.tableApply function passes the ore.frame table proxy to the input function as the dat argument. The input function creates a model, which the ore.tableApply function returns as an ore.object object.
Example 6-11 Using the ore.tableApply Function
library(e1071)
nbmod <- ore.tableApply(
ore.push(iris),
function(dat) {
library(e1071)
dat$Species <- as.factor(dat$Species)
naiveBayes(Species ~ ., dat)
})
class(nbmod)
nbmod
Listing for Example 6-11
R> nbmod <- ore.tableApply(
+ ore.push(iris),
+ function(dat) {
+ library(e1071)
+ dat$Species <- as.factor(dat$Species)
+ naiveBayes(Species ~ ., dat)
+ })
R> class(nbmod)
[1] "ore.object"
attr(,"package")
[1] "OREembed"
R> nbmod
Naive Bayes Classifier for Discrete Predictors
Call:
naiveBayes.default(x = X, y = Y, laplace = laplace)
A-priori probabilities:
Y
setosa versicolor virginica
0.3333333 0.3333333 0.3333333
Conditional probabilities:
Sepal.Length
Y [,1] [,2]
setosa 5.006 0.3524897
versicolor 5.936 0.5161711
virginica 6.588 0.6358796
Sepal.Width
Y [,1] [,2]
setosa 3.428 0.3790644
versicolor 2.770 0.3137983
virginica 2.974 0.3224966
Petal.Length
Y [,1] [,2]
setosa 1.462 0.1736640
versicolor 4.260 0.4699110
virginica 5.552 0.5518947
Petal.Width
Y [,1] [,2]
setosa 0.246 0.1053856
versicolor 1.326 0.1977527
virginica 2.026 0.2746501
Using the ore.groupApply Function
The ore.groupApply function invokes an R script with an ore.frame as the input data. The ore.groupApply function passes the ore.frame to the user-defined input function as the first argument to that function. The INDEX argument to the ore.groupApply function specifies the name of a column of the ore.frame by which Oracle Database partitions the rows for processing by the user-defined R function. The ore.groupApply function can use data-parallel execution, in which one or more R engines perform the same R function, or task, on different partitions of data.
The syntax of the ore.groupApply function is the following:
ore.groupApply(X, INDEX, FUN, ..., FUN.VALUE = NULL, FUN.NAME = NULL,
parallel = getOption("ore.parallel", NULL))
The ore.groupApply function returns an ore.list object or an ore.frame object.
Examples of the use of the ore.groupApply function are in the following topics:
See Also:
-
"Arguments for Functions that Run Scripts" for descriptions of the arguments to function
ore.groupApply -
"Installing a Third-Party Package for Use in Embedded R Execution"
Partitioning on a Single Column
Example 6-12 uses the C50 package, which has functions that build decision tree and rule-based models. The package also provides training and testing data sets. Example 6-12 builds C5.0 models on the churnTrain training data set from the churn data set of the C50 package, with the goal of building one churn model on the data for each state. The example does the following:
-
Loads the
C50package and then thechurndata set. -
Uses the
ore.createfunction to create theCHURN_TRAINdatabase table and its proxyore.frameobject fromchurnTrain, adata.frameobject. -
Specifies
CHURN_TRAIN, the proxyore.frameobject, as the first argument to theore.groupApplyfunction and specifies thestatecolumn as theINDEXargument. Theore.groupApplyfunction partitions the data on thestatecolumn and invokes the user-defined function on each partition. -
Creates the variable
modList, which gets theore.listobject returned by theore.groupApplyfunction. Theore.listobject contains the results from the execution of the user-defined function on each partition of the data. In this case, it is one C5.0 model per state, with each model stored as anore.objectobject. -
Specifies the user-defined function. The first argument of the user-defined function receives one partition of the data, which in this case is all of the data associated with a single state.
The user-defined function does the following:
-
Loads the C50 package so that it is available to the function when it executes in an R engine in the database.
-
Deletes the
statecolumn from thedata.frameso that the column is not included in the model. -
Converts the columns to factors because, although the
ore.framedefined factors, when they are loaded to the user-defined function, factors appear as character vectors. -
Builds a model for a state and returns it.
-
-
Uses the
ore.pullfunction to retrieve the model from the database as themod.MAvariable and then invokes thesummaryfunction on it. The class ofmod.MAisC5.0.
Example 6-12 Using the ore.groupApply Function
library(C50)
data("churn")
ore.create(churnTrain, "CHURN_TRAIN")
modList <- ore.groupApply(
CHURN_TRAIN,
INDEX=CHURN_TRAIN$state,
function(dat) {
library(C50)
dat$state <- NULL
dat$churn <- as.factor(dat$churn)
dat$area_code <- as.factor(dat$area_code)
dat$international_plan <- as.factor(dat$international_plan)
dat$voice_mail_plan <- as.factor(dat$voice_mail_plan)
C5.0(churn ~ ., data = dat, rules = TRUE)
});
mod.MA <- ore.pull(modList$MA)
summary(mod.MA)
Listing for Example 6-12
R> library(C50)
R> data(churn)
R>
R> ore.create(churnTrain, "CHURN_TRAIN")
R>
R> modList <- ore.groupApply(
+ CHURN_TRAIN,
+ INDEX=CHURN_TRAIN$state,
+ function(dat) {
+ library(C50)
+ dat$state <- NULL
+ dat$churn <- as.factor(dat$churn)
+ dat$area_code <- as.factor(dat$area_code)
+ dat$international_plan <- as.factor(dat$international_plan)
+ dat$voice_mail_plan <- as.factor(dat$voice_mail_plan)
+ C5.0(churn ~ ., data = dat, rules = TRUE)
+ });
R> mod.MA <- ore.pull(modList$MA)
R> summary(mod.MA)
Call:
C5.0.formula(formula = churn ~ ., data = dat, rules = TRUE)
C5.0 [Release 2.07 GPL Edition] Thu Feb 13 15:09:10 2014
-------------------------------
Class specified by attribute `outcome'
Read 65 cases (19 attributes) from undefined.data
Rules:
Rule 1: (52/1, lift 1.2)
international_plan = no
total_day_charge <= 43.04
-> class no [0.963]
Rule 2: (5, lift 5.1)
total_day_charge > 43.04
-> class yes [0.857]
Rule 3: (6/1, lift 4.4)
area_code in {area_code_408, area_code_415}
international_plan = yes
-> class yes [0.750]
Default class: no
Evaluation on training data (65 cases):
Rules
----------------
No Errors
3 2( 3.1%) <<
(a) (b) <-classified as
---- ----
53 1 (a): class no
1 10 (b): class yes
Attribute usage:
89.23% international_plan
87.69% total_day_charge
9.23% area_code
Time: 0.0 secs
Partitioning on Multiple Columns
The ore.groupApply function takes only a single column for the INDEX argument; however, you can create a new column that is the concatenation of the columns you want to use and provide this new column as the INDEX argument.
Example 6-13 uses data from the CHURN_TRAIN data set to build an rpart model that produces rules on the partitions of data specified, which are the voice_mail_plan and international_plan columns. The example uses the R table function to show the number of rows to expect in each partition. It then adds a new column that pastes together the two columns of interest to create a new column named vmp_ip.g1
The example next invokes the ore.scriptDrop function to ensure that no script by the specified name exists in the R script repository. It then uses the ore.scriptCreate function to define a script named my.rpartFunction and to store it in the repository. The stored script defines a function that takes a data source and a prefix to use for naming Oracle R Enterprise datastore objects. Each invocation of the function my.rpartFunction receives data from one of the partitions identified in vmp_ip. Because the source partition columns are constants, the function sets them to NULL. It converts the character vectors to factors, builds a model to predict churn, and saves it in an appropriately named datastore. The function creates a list to return the specific partition column values, the distribution of churn values, and the model itself.
The example then loads the rpart library, sets the datastore prefix, and invokes ore.groupApply using the derived column vmp_ip as the value of the INDEX argument and my.rpartFunction as the value of the FUN.NAME argument to invoke the user-defined function stored in the R script repository. The ore.groupApply function uses an optional argument to pass the datastorePrefix variable to the user-defined function. It uses the optional argument ore.connect to connect to the database when executing the user-defined function. The ore.groupApply function returns an ore.list object as the variable res.
The example displays the first entry in the list returned. It then invokes the ore.load function to load the model for the case where the customer has both the voice mail plan and the international plan.
Example 6-13 Using ore.groupApply for Partitioning Data on Multiple Columns
library(C50)
data(churn)
ore.drop("CHURN_TRAIN")
ore.create(churnTrain, "CHURN_TRAIN")
table(CHURN_TRAIN$international_plan, CHURN_TRAIN$voice_mail_plan)
CT <- CHURN_TRAIN
CT$vmp_ip <- paste(CT$voice_mail_plan, CT$international_plan,sep = "-")
options(width = 80)
head(CT, 3)
ore.scriptDrop("my.rpartFunction")
ore.scriptCreate("my.rpartFunction",
function(dat, datastorePrefix) {
library(rpart)
vmp <- dat[1, "voice_mail_plan"]
ip <- dat[1, "international_plan"]
datastoreName <- paste(datastorePrefix, vmp, ip, sep = "_")
dat$voice_mail_plan <- NULL
dat$international_plan <- NULL
dat$state <- as.factor(dat$state)
dat$churn <- as.factor(dat$churn)
dat$area_code <- as.factor(dat$area_code)
mod <- rpart(churn ~ ., data = dat)
ore.save(mod, name = datastoreName, overwrite = TRUE)
list(voice_mail_plan = vmp,
international_plan = ip,
churn.table = table(dat$churn),
rpart.model = mod)
})
library(rpart)
datastorePrefix = "my.rpartModel"
res <- ore.groupApply(CT, INDEX = CT$vmp_ip,
FUN.NAME = "my.rpartFunction",
datastorePrefix = datastorePrefix,
ore.connect = TRUE)
res[[1]]
ore.load(name=paste(datastorePrefix, "yes", "yes", sep = "_"))
mod
Listing for Example 6-13
R> library(C50)
R> data(churn)
R> ore.drop("CHURN_TRAIN")
R> ore.create(churnTrain, "CHURN_TRAIN")
R>
R> table(CHURN_TRAIN$international_plan, CHURN_TRAIN$voice_mail_plan)
no yes
no 2180 830
yes 231 92
R> CT <- CHURN_TRAIN
R> CT$vmp_ip <- paste(CT$voice_mail_plan, CT$international_plan, sep = "-")
R> options(width = 80)
R> head(CT, 3)
state account_length area_code international_plan voice_mail_plan
1 KS 128 area_code_415 no yes
2 OH 107 area_code_415 no yes
3 NJ 137 area_code_415 no no
number_vmail_messages total_day_minutes total_day_calls total_day_charge
1 25 265.1 110 45.07
2 26 161.6 123 27.47
3 0 243.4 114 41.38
total_eve_minutes total_eve_calls total_eve_charge total_night_minutes
1 197.4 99 16.78 244.7
2 195.5 103 16.62 254.4
3 121.2 110 10.30 162.6
total_night_calls total_night_charge total_intl_minutes total_intl_calls
1 91 11.01 10.0 3
2 103 11.45 13.7 3
3 104 7.32 12.2 5
total_intl_charge number_customer_service_calls churn vmp_ip
1 2.70 1 no yes-no
2 3.70 1 no yes-no
3 3.29 0 no no-no
R>
R> ore.scriptDrop("my.rpartFunction")
R> ore.scriptCreate("my.rpartFunction",
+ function(dat, datastorePrefix) {
+ library(rpart)
+ vmp <- dat[1, "voice_mail_plan"]
+ ip <- dat[1, "international_plan"]
+ datastoreName <- paste(datastorePrefix, vmp, ip, sep = "_")
+ dat$voice_mail_plan <- NULL
+ dat$international_plan <- NULL
+ dat$state <- as.factor(dat$state)
+ dat$churn <- as.factor(dat$churn)
+ dat$area_code <- as.factor(dat$area_code)
+ mod <- rpart(churn ~ ., data = dat)
+ ore.save(mod, name = datastoreName, overwrite = TRUE)
+ list(voice_mail_plan = vmp,
+ international_plan = ip,
+ churn.table = table(dat$churn),
+ rpart.model = mod)
+ })
R>
R> library(rpart)
R> datastorePrefix = "my.rpartModel"
R>
R> res <- ore.groupApply(CT, INDEX = CT$vmp_ip,
+ FUN.NAME = "my.rpartFunction",
+ datastorePrefix = datastorePrefix,
+ ore.connect = TRUE)
R> res[[1]]
$voice_mail_plan
[1] "no"
$international_plan
[1] "no"
$churn.table
no yes
1878 302
$rpart.model
n= 2180
node), split, n, loss, yval, (yprob)
* denotes terminal node
1) root 2180 302 no (0.86146789 0.13853211)
2) total_day_minutes< 263.55 2040 192 no (0.90588235 0.09411765)
4) number_customer_service_calls< 3.5 1876 108 no (0.94243070 0.05756930)
8) total_day_minutes< 223.25 1599 44 no (0.97248280 0.02751720) *
9) total_day_minutes>=223.25 277 64 no (0.76895307 0.23104693)
18) total_eve_minutes< 242.35 210 18 no (0.91428571 0.08571429) *
19) total_eve_minutes>=242.35 67 21 yes (0.31343284 0.68656716)
38) total_night_minutes< 174.2 17 4 no (0.76470588 0.23529412) *
39) total_night_minutes>=174.2 50 8 yes (0.16000000 0.84000000) *
5) number_customer_service_calls>=3.5 164 80 yes (0.48780488 0.51219512)
10) total_day_minutes>=160.2 95 22 no (0.76842105 0.23157895)
20) state=AL,AZ,CA,CO,DC,DE,FL,HI,KS,KY,MA,MD,ME,MI,NC,ND,NE,NH,NM,OK,OR,SC,TN,VA,VT,WY 56 2 no (0.96428571 0.03571429) *
21) state=AK,AR,CT,GA,IA,ID,MN,MO,NJ,NV,NY,OH,RI,TX,UT,WA,WV 39 19 yes (0.48717949 0.51282051)
42) total_day_minutes>=182.3 21 5 no (0.76190476 0.23809524) *
43) total_day_minutes< 182.3 18 3 yes (0.16666667 0.83333333) *
11) total_day_minutes< 160.2 69 7 yes (0.10144928 0.89855072) *
3) total_day_minutes>=263.55 140 30 yes (0.21428571 0.78571429)
6) total_eve_minutes< 167.3 29 7 no (0.75862069 0.24137931)
12) state=AK,AR,AZ,CO,CT,FL,HI,IN,KS,LA,MD,ND,NM,NY,OH,UT,WA,WV 21 0 no (1.00000000 0.00000000) *
13) state=IA,MA,MN,PA,SD,TX,WI 8 1 yes (0.12500000 0.87500000) *
7) total_eve_minutes>=167.3 111 8 yes (0.07207207 0.92792793) *
R> ore.load(name = paste(datastorePrefix, "yes", "yes", sep = "_"))
[1] "mod"
R> mod
n= 92
node), split, n, loss, yval, (yprob)
* denotes terminal node
1) root 92 36 no (0.60869565 0.39130435)
2) total_intl_minutes< 13.1 71 15 no (0.78873239 0.21126761)
4) total_intl_calls>=2.5 60 4 no (0.93333333 0.06666667)
8) state=AK,AR,AZ,CO,CT,DC,DE,FL,GA,HI,ID,IL,IN,KS,MD,MI,MO,MS,MT,NC,ND,NE,NH,NJ,OH,SC,SD,UT,VA,WA,WV,WY 53 0 no (1.00000000 0.00000000) *
9) state=ME,NM,VT,WI 7 3 yes (0.42857143 0.57142857) *
5) total_intl_calls< 2.5 11 0 yes (0.00000000 1.00000000) *
3) total_intl_minutes>=13.1 21 0 yes (0.00000000 1.00000000) *
Using the ore.rowApply Function
The ore.rowApply function invokes an R script with an ore.frame as the input data. The ore.rowApply function passes the ore.frame to the user-defined input function as the first argument to that function. The rows argument to the ore.rowApply function specifies the number of rows to pass to each invocation of the user-defined R function. The last chunk or rows may have fewer rows than the number specified. The ore.rowApply function can use data-parallel execution, in which one or more R engines perform the same R function, or task, on different partitions of data.
The syntax of the ore.rowApply function is the following:
ore.rowApply(X, FUN, ..., FUN.VALUE = NULL, FUN.NAME = NULL, rows = 1,
parallel = getOption("ore.parallel", NULL))
The ore.rowApply function returns an ore.list object or an ore.frame object.
See Also:
-
"Arguments for Functions that Run Scripts" for descriptions of the arguments to function
ore.rowApply -
"Installing a Third-Party Package for Use in Embedded R Execution"
Example 6-14 uses the e1071 package, previously downloaded from CRAN. The example also uses the nbmod object, which is the Naive Bayes model created in Example 6-11, "Using the ore.tableApply Function".
Example 6-14 does the following:
-
Loads the package
e1071. -
Pushes the
irisdata set to the database as theIRIStemporary table andore.frameobject. -
Creates a copy of
IRISasIRIS_PREDand adds the PRED column toIRIS_PREDto contain the predictions. -
Invokes the
ore.rowApplyfunction, passing theIRISore.frameas the data source for user-defined R function and the user-defined R function itself. -
The user-defined function does the following:
-
Loads the package
e1071so that it is available to the R engine or engines that run in the database. -
Converts the Species column to a factor because, although the
ore.framedefined factors, when they are loaded to the user-defined function, factors appear as character vectors. -
Invokes the
predictmethod and returns theresobject, which contains the predictions in the column added to the data set.
-
-
The example pulls the model to the client R session.
-
Passes
IRIS_PREDas the argumentFUN.VALUE, which specifies the structure of the object that theore.rowApplyfunction returns. -
Specifies the number of rows to pass to each invocation of the user-defined function.
-
Displays the class of
res, and invokes thetablefunction to display the Species column and the PRED column of theresobject.
Example 6-14 Using the ore.rowApply Function
library(e1071)
IRIS <- ore.push(iris)
IRIS_PRED <- IRIS
IRIS_PRED$PRED <- "A"
res <- ore.rowApply(
IRIS,
function(dat, nbmod) {
library(e1071)
dat$Species <- as.factor(dat$Species)
dat$PRED <- predict(nbmod, newdata = dat)
dat
},
nbmod = ore.pull(nbmod),
FUN.VALUE = IRIS_PRED,
rows = 10)
class(res)
table(res$Species, res$PRED)
Listing for Example 6-14
R> library(e1071)
R> IRIS <- ore.push(iris)
R> IRIS_PRED <- IRIS
R> IRIS_PRED$PRED <- "A"
R> res <- ore.rowApply(
+ IRIS ,
+ function(dat, nbmod) {
+ library(e1071)
+ dat$Species <- as.factor(dat$Species)
+ dat$PRED <- predict(nbmod, newdata = dat)
+ dat
+ },
+ nbmod = ore.pull(nbmod),
+ FUN.VALUE = IRIS_PRED,
+ rows = 10)
R> class(res)
[1] "ore.frame"
attr(,"package")
[1] "OREbase"
R> table(res$Species, res$PRED)
setosa versicolor virginica
setosa 50 0 0
versicolor 0 47 3
virginica 0 3 47
As Example 6-12 does, Example 6-15 uses the C50 package to score churn data (that is, to predict which customers are likely to churn) using C5.0 models. However, instead of partitioning the data by a column, Example 6-15 partitions the data by a number of rows. The example scores the customers from the specified state in parallel. The example uses datastores and saves functions to the R script repository, which allows the functions to be used by the Oracle R Enterprise SQL API functions.
Example 6-15 first loads C50 package and the data sets. The example deletes the datastores with names containing myC5.0modelFL, if they exist. It invokes ore.drop to delete the CHURN_TEST table, if it exists, and then invokes ore.create to create the CHURN_TEST table from the churnTest data set.
The example next invokes ore.getLevels, which returns a list of the levels for each factor column. The invocation excludes the first column, which is state, because the levels for that column are not needed. Getting the levels first can ensure that all possible levels are provided during model building, even if some rows do not have values for some of the levels. The ore.delete invocation ensures that no datastore with the specified name exists and the ore.save invocation saves the xlevels object in the datastore named myXLevels.
Example 6-15 creates a user-defined function, myC5.0FunctionForLevels, that generates a C5.0 model. The function uses the list of levels returned by function ore.getXlevels instead of computing the levels using the as.factor function as the user-defined function does in Example 6-12. It uses the levels to convert the column type from character vector to factor. The function myC5.0FunctionForLevels returns the value TRUE. The example saves the function in the R script repository.
The example next gets a list of datastores that have names that include the specified string and deletes those datastores if they exist.
The example then invokes ore.groupApply, which invokes function myC5.0FunctionForLevels on each state in the CHURN_TEST data. To each myC5.0FunctionForLevels invocation, ore.groupApply passes the datastore that contains the xlevels object and a prefix to use in naming the datastore generated by myC5.0FunctionForLevels. It also passes the ore.connect control argument to connect to the database in the embedded R function, which enables the use of objects stored in a datastore. The ore.groupApply invocation returns a list that contains the results of all of the invocations of myC5.0FunctionForLevels.
The example pulls the result over to the local R session and verifies that myC5.0FunctionForLevels returned TRUE for each state in the data source.
Example 6-15 next creates another user-defined another function, myScoringFunction, and stores it in the R script repository. The function scores a C5.0 model for the levels of a state and returns the results in a data.frame.
The example then invokes function ore.rowApply. It filters the input data to use only data for the state of Massachusetts. It specifies myScoringFunction as the function to invoke and passes that user-defined function the name of the datastore that contains the xlevels object and a prefix to use in loading the datastore that contains the C5.0 model for the state. The ore.rowApply invocation specifies invoking myScoringFunction on 200 rows of the data set in each parallel R engine. It uses the FUN.VALUE argument so that ore.rowApply returns an ore.frame that contains the results of all of the myScoringFunction invocations. The variable scores gets the results of the ore.rowApply invocation.
Finally, Example 6-15 prints the scores object and then uses the table function to display the confusion matrix for the scoring.
See Also:
Example 6-24, "Using an rqRowEval Function" for an invocation of therqRowEval function that produces the same result as the ore.rowApply function in Example 6-15Example 6-15 Using the ore.rowApply Function with Datastores and Scripts
library(C50)
data(churn)
ore.drop("CHURN_TEST"
ore.create(churnTest, "CHURN_TEST")
xlevels <- ore.getXlevels(~ ., CHURN_TEST[,-1])
ore.delete("myXLevels")
ore.save(xlevels, name = "myXLevels")
ore.scriptDrop("myC5.0FunctionForLevels")
ore.scriptCreate("myC5.0FunctionForLevels",
function(dat, xlevelsDatastore, datastorePrefix) {
library(C50)
state <- dat[1,"state"]
datastoreName <- paste(datastorePrefix, dat[1, "state"], sep = "_")
dat$state <- NULL
ore.load(name = xlevelsDatastore)
for (j in names(xlevels))
dat[[j]] <- factor(dat[[j]], levels = xlevels[[j]])
c5mod <- C5.0(churn ~ ., data = dat, rules = TRUE)
ore.save(c5mod, name = datastoreName)
TRUE
})
ds.v <- ore.datastore(pattern= "myC5.0modelFL")$datastore.name
for (ds in ds.v) ore.delete(name = ds)
res <- ore.groupApply(CHURN_TEST,
INDEX=CHURN_TEST$state,
FUN.NAME = "myC5.0FunctionForLevels",
xlevelsDatastore = "myXLevels",
datastorePrefix = "myC5.0modelFL",
ore.connect = TRUE)
res <- ore.pull(res)
all(as.logical(res) == TRUE)
ore.scriptDrop("myScoringFunction")
ore.scriptCreate("myScoringFunction",
function(dat, xlevelsDatastore, datastorePrefix) {
library(C50)
state <- dat[1,"state"]
datastoreName <- paste(datastorePrefix,state,sep="_")
dat$state <- NULL
ore.load(name = xlevelsDatastore)
for (j in names(xlevels))
dat[[j]] <- factor(dat[[j]], levels = xlevels[[j]])
ore.load(name = datastoreName)
res <- data.frame(pred = predict(c5mod, dat, type = "class"),
actual = dat$churn,
state = state)
res
}
)
scores <- ore.rowApply(
CHURN_TEST[CHURN_TEST$state == "MA",],
FUN.NAME = "myScoringFunction",
xlevelsDatastore = "myXLevels",
datastorePrefix = "myC5.0modelFL",
ore.connect = TRUE, parallel = TRUE,
FUN.VALUE = data.frame(pred = character(0),
actual = character(0),
state = character(0)),
rows=200)
scores
table(scores$actual, scores$pred)
Listing for Example 6-15
R> library(C50)
R> data(churn)
R>
R> ore.drop("CHURN_TEST"
R> ore.create(churnTest, "CHURN_TEST")
R>
R> xlevels <- ore.getXlevels(~ ., CHURN_TEST[,-1])
R> ore.delete("myXLevels")
[1] "myXLevels
R> ore.save(xlevels, name = "myXLevels")
R>
R> ore.scriptDrop("myC5.0FunctionForLevels")
R> ore.scriptCreate("myC5.0FunctionForLevels",
+ function(dat, xlevelsDatastore, datastorePrefix) {
+ library(C50)
+ state <- dat[1,"state"]
+ datastoreName <- paste(datastorePrefix, dat[1, "state"], sep = "_")
+ dat$state <- NULL
+ ore.load(name = xlevelsDatastore)
+ for (j in names(xlevels))
+ dat[[j]] <- factor(dat[[j]], levels = xlevels[[j]])
+ c5mod <- C5.0(churn ~ ., data = dat, rules = TRUE)
+ ore.save(c5mod, name = datastoreName)
+ TRUE
+ })
R>
R> ds.v <- ore.datastore(pattern="myC5.0modelFL")$datastore.name
R> for (ds in ds.v) ore.delete(name=ds)
R>
R> res <- ore.groupApply(CHURN_TEST,
+ INDEX=CHURN_TEST$state,
+ FUN.NAME="myC5.0FunctionForLevels",
+ xlevelsDatastore = "myXLevels",
+ datastorePrefix = "myC5.0modelFL",
+ ore.connect = TRUE)
R> res <- ore.pull(res)
R> all(as.logical(res) == TRUE)
[1] TRUE
R>
R> ore.scriptDrop("myScoringFunction")
R> ore.scriptCreate("myScoringFunction",
+ function(dat, xlevelsDatastore, datastorePrefix) {
+ library(C50)
+ state <- dat[1,"state"]
+ datastoreName <- paste(datastorePrefix,state,sep="_")
+ dat$state <- NULL
+ ore.load(name = xlevelsDatastore)
+ for (j in names(xlevels))
+ dat[[j]] <- factor(dat[[j]], levels = xlevels[[j]])
+ ore.load(name = datastoreName)
+ res <- data.frame(pred = predict(c5mod, dat, type="class"),
+ actual = dat$churn,
+ state = state)
+ res
+ }
+ )
R>
R> scores <- ore.rowApply(
+ CHURN_TEST[CHURN_TEST$state =="MA",],
+ FUN.NAME = "myScoringFunction",
+ xlevelsDatastore = "myXLevels",
+ datastorePrefix = "myC5.0modelFL",
+ ore.connect = TRUE, parallel = TRUE,
+ FUN.VALUE = data.frame(pred=character(0),
+ actual=character(0),
+ state=character(0)),
+ rows=200
R>
R> scores
pred actual state
1 no no MA
2 no no MA
3 no no MA
4 no no MA
5 no no MA
6 no yes MA
7 yes yes MA
8 yes yes MA
9 no no MA
10 no no MA
11 no no MA
12 no no MA
13 no no MA
14 no no MA
15 yes yes MA
16 no no MA
17 no no MA
18 no no MA
19 no no MA
20 no no MA
21 no no MA
22 no no MA
23 no no MA
24 no no MA
25 no no MA
26 no no MA
27 no no MA
28 no no MA
29 no yes MA
30 no no MA
31 no no MA
32 no no MA
33 yes yes MA
34 no no MA
35 no no MA
36 no no MA
37 no no MA
38 no no MA
Warning message:
ORE object has no unique key - using random order
R> table(scores$actual, scores$pred)
no yes
no 32 0
yes 2 4
Using the ore.indexApply Function
The ore.indexApply function executes the specified user-defined input function using data that is generated by the input function. It supports task-parallel execution, in which one or more R engines perform the same or different calculations, or task. The times argument to the ore.indexApply function specifies the number of times that the input function executes in the database. Any required data must be explicitly generated or loaded within the input function.
The syntax of the ore.indexApply function is the following:
ore.indexApply(times, FUN, ..., FUN.VALUE = NULL, FUN.NAME = NULL,
parallel = getOption("ore.parallel", NULL))
The ore.indexApply function returns an ore.list object or an ore.frame object.
Examples of the use of the ore.indexApply function are in the following topics:
See Also:
-
"Arguments for Functions that Run Scripts" for descriptions of the arguments to function
ore.indexApply -
"Installing a Third-Party Package for Use in Embedded R Execution"
Simple Example of Using the ore.indexApply Function
Example 6-16 invokes ore.indexApply and specifies that it execute the input function five times in parallel. It displays the class of the result, which is ore.list, and then displays the result.
Example 6-16 Using the ore.indexApply Function
res <- ore.indexApply(5,
function(index) {
paste("IndexApply:", index)
},
parallel = TRUE)
class(res)
res
Listing for Example 6-16
R> res <- ore.indexApply(5,
+ function(index) {
+ paste("IndexApply:", index)
+ },
+ parallel = TRUE)
R> class(res)
[1] "ore.list"
attr(,"package")
[1] "OREembed"
R> res
$`1`
[1] "IndexApply: 1"
$`2`
[1] "IndexApply: 2"
$`3`
[1] "IndexApply: 3"
$`4`
[1] "IndexApply: 4"
$`5`
[1] "IndexApply: 5"
Column-Parallel Use Case
Example 6-17 uses the R summary function to compute in parallel summary statistics on the first four numeric columns of the iris data set. The example combines the computations into a final result. The first argument to the ore.indexApply function is 4, which specifies the number of columns to summarize in parallel. The user-defined input function takes one argument, index, which will be a value between 1 and 4 and which specifies the column to summarize.
The example invokes the summary function on the specified column. The summary invocation returns a single row, which contains the summary statistics for the column. The example converts the result of the summary invocation into a data.frame and adds the column name to it.
The example next uses the FUN.VALUE argument to the ore.indexApply function to define the structure of the result of the function. The result is then returned as an ore.frame object with that structure.
Example 6-17 Using the ore.indexApply Function and Combining Results
res <- NULL
res <- ore.indexApply(4,
function(index) {
ss <- summary(iris[, index])
attr.names <- attr(ss, "names")
stats <- data.frame(matrix(ss, 1, length(ss)))
names(stats) <- attr.names
stats$col <- names(iris)[index]
stats
},
FUN.VALUE=data.frame(Min. = numeric(0),
"1st Qu." = numeric(0),
Median = numeric(0),
Mean = numeric(0),
"3rd Qu." = numeric(0),
Max. = numeric(0),
Col = character(0)),
parallel = TRUE)
res
Listing for Example 6-17
R> res <- NULL
R> res <- ore.indexApply(4,
+ function(index) {
+ ss <- summary(iris[, index])
+ attr.names <- attr(ss, "names")
+ stats <- data.frame(matrix(ss, 1, length(ss)))
+ names(stats) <- attr.names
+ stats$col <- names(iris)[index]
+ stats
+ },
+ FUN.VALUE=data.frame(Min. = numeric(0),
+ "1st Qu." = numeric(0),
+ Median = numeric(0),
+ Mean = numeric(0),
+ "3rd Qu." = numeric(0),
+ Max. = numeric(0),
+ Col = character(0)),
+ parallel = TRUE)
R> res
Min. X1st.Qu. Median Mean X3rd.Qu. Max. Col
1 2.0 2.8 3.00 3.057 3.3 4.4 Sepal.Width
2 4.3 5.1 5.80 5.843 6.4 7.9 Sepal.Length
3 0.1 0.3 1.30 1.199 1.8 2.5 Petal.Width
4 1.0 1.6 4.35 3.758 5.1 6.9 Petal.Length
Warning message:
ORE object has no unique key - using random order
Simulations Use Case
You can use the ore.indexApply function in simulations, which can take advantage of high-performance computing hardware like an Oracle Exadata Database Machine. Example 6-18 takes multiple samples from a random normal distribution to compare the distribution of the summary statistics. Each simulation occurs in a separate R engine in the database, in parallel, up to the degree of parallelism allowed by the database.
Example 6-18 defines variables for the sample size, the mean and standard deviations of the random numbers, and the number of simulations to perform. The example specifies num.simulations as the first argument to the ore.indexApply function. The ore.indexApply function passes num.simulations to the user-defined function as the index argument. This input function then sets the random seed based on the index so that each invocation of the input function generates a different set of random numbers.
The input function next uses the rnorm function to produce sample.size random normal values. It invokes the summary function on the vector of random numbers, and then prepares a data.frame as the result it returns. The ore.indexApply function specifies the FUN.VALUE argument so that it returns an ore.frame that structures the combined results of the simulations. The res variable gets the ore.frame returned by the ore.indexApply function.
To get the distribution of samples, the example invokes the boxplot function on the data.frame that is the result of using the ore.pull function to bring selected columns from res to the client.
Example 6-18 Using the ore.indexApply Function in a Simulation
res <- NULL
sample.size = 1000
mean.val = 100
std.dev.val = 10
num.simulations = 1000
res <- ore.indexApply(num.simulations,
function(index, sample.size = 1000, mean = 0, std.dev = 1) {
set.seed(index)
x <- rnorm(sample.size, mean, std.dev)
ss <- summary(x)
attr.names <- attr(ss, "names")
stats <- data.frame(matrix(ss, 1, length(ss)))
names(stats) <- attr.names
stats$index <- index
stats
},
FUN.VALUE=data.frame(Min. = numeric(0),
"1st Qu." = numeric(0),
Median = numeric(0),
Mean = numeric(0),
"3rd Qu." = numeric(0),
Max. = numeric(0),
Index = numeric(0)),
parallel = TRUE,
sample.size = sample.size,
mean = mean.val, std.dev = std.dev.val)
options("ore.warn.order" = FALSE)
head(res, 3)
tail(res, 3)
boxplot(ore.pull(res[, 1:6]),
main=sprintf("Boxplot of %d rnorm samples size %d, mean=%d, sd=%d",
num.simulations, sample.size, mean.val, std.dev.val))
Listing for Example 6-18
R> res <- ore.indexApply(num.simulations,
+ function(index, sample.size = 1000, mean = 0, std.dev = 1) {
+ set.seed(index)
+ x <- rnorm(sample.size, mean, std.dev)
+ ss <- summary(x)
+ attr.names <- attr(ss, "names")
+ stats <- data.frame(matrix(ss, 1, length(ss)))
+ names(stats) <- attr.names
+ stats$index <- index
+ stats
+ },
+ FUN.VALUE=data.frame(Min. = numeric(0),
+ "1st Qu." = numeric(0),
+ Median = numeric(0),
+ Mean = numeric(0),
+ "3rd Qu." = numeric(0),
+ Max. = numeric(0),
+ Index = numeric(0)),
+ parallel = TRUE,
+ sample.size = sample.size,
+ mean = mean.val, std.dev = std.dev.val)
R> options("ore.warn.order" = FALSE)
R> head(res, 3)
Min. X1st.Qu. Median Mean X3rd.Qu. Max. Index
1 67.56 93.11 99.42 99.30 105.8 128.0 847
2 67.73 94.19 99.86 100.10 106.3 130.7 258
3 65.58 93.15 99.78 99.82 106.2 134.3 264
R> tail(res, 3)
Min. X1st.Qu. Median Mean X3rd.Qu. Max. Index
1 65.02 93.44 100.2 100.20 106.9 134.0 5
2 71.60 93.34 99.6 99.66 106.4 131.7 4
3 69.44 93.15 100.3 100.10 106.8 135.2 3
R> boxplot(ore.pull(res[, 1:6]),
+ main=sprintf("Boxplot of %d rnorm samples size %d, mean=%d, sd=%d",
+ num.simulations, sample.size, mean.val, std.dev.val))
Figure 6-2 Display of the boxplot Function in Example 6-18
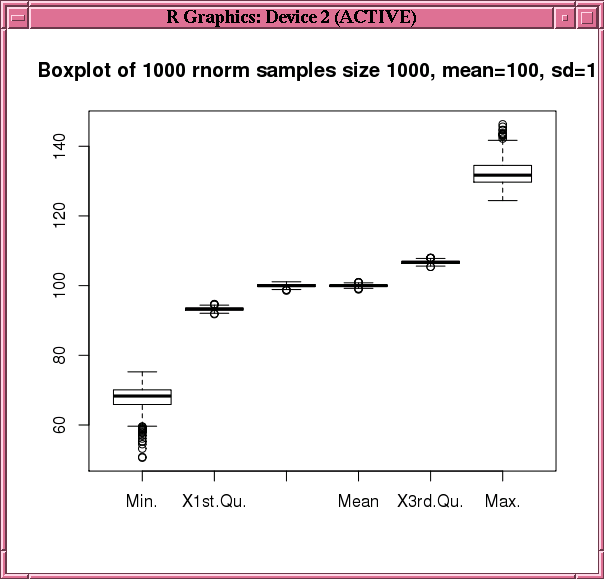
Description of "Figure 6-2 Display of the boxplot Function in Example 6-18"
SQL Interface for Embedded R Execution
The SQL interface for Oracle R Enterprise embedded R execution allows you to execute R scripts in production database applications. The SQL interface has functions for adding and removing a script from the R script repository, for executing an R script, and for deleting an Oracle R Enterprise datastore.
This SQL interface is described in the following topics:
About Oracle R Enterprise SQL Table Functions
Oracle R Enterprise provides SQL table functions that are equivalents of most of the R interface functions for embedded R execution. Executing a SELECT FROM TABLE statement and specifying one of the table functions results in the invocation of the specified R script. The script runs in one or more R engines on the Oracle Database server.
The SQL table functions for embedded R execution are:
-
rqEval -
rqGroupEval
-
rqRowEval -
rqTableEval
The R interface functions and the SQL equivalents are listed in Table 6-1.
For the rqGroupEval function, Oracle R Enterprise provides a generic implementation of the group apply functionality in SQL. You must write a table function that captures the structure of the input cursor.
Some general aspects of the SQL table functions are described in the following topics in this section:
For more information about the functions, including examples of their use, see the following:
Parameters of the SQL Table Functions
The SQL table functions have some parameters in common and some functions have parameters that are unique to that function. The parameters of the SQL table functions are the following.
Table 6-2 SQL Table Function Parameters
| Parameter | Description |
|---|---|
|
|
A cursor that specifies the data that is input to the R function specified by |
|
|
A cursor that specifies arguments to pass to the R function. The parameters cursor consists of a single row of scalar values. An argument can be a string or a numeric value. You can specify multiple arguments in the cursor. Arguments to an R function are case sensitive, so you should put names, such as a column name, in double quotes. In the cursor, you can also specify as scalar values an Oracle R Enterprise control argument or the names of serialized R objects, such as predictive models, that are in an Oracle R Enterprise datastore. The value of this parameters cursor can be |
|
|
An output table definition. The value of this argument can be |
|
|
For the rqGroupEval function, the name of the grouping column. |
|
|
For the |
|
|
The name that identifies an R function in the R script repository. The |
Return Value of SQL Table Functions
The Oracle R Enterprise SQL table functions return a table. The structure and contents of the table are determined by the results of the R function passed to the SQL table function and by the OUT_QRY parameter. The R function can return a data.frame object, other R objects, and graphics. The structure of the table that represents the results of the R function is specified by one of the following OUT_QRY values:
-
NULL, which results in a table that has a serialized object that can contain both data and image objects. -
A table signature specified in a
SELECTstatement, which results in a table that has the defined structure. The result of the R function must be adata.frame. No images are returned. -
The string
'XML', which results in a table that has a CLOB that can contain both structured data and graph images in an XML string. The non-image R objects, such asdata.frameormodelobjects, are provided first, followed by the base 64 encoding of a PNG representation of the image. -
The string
'PNG', which results in a table that has a BLOB that contains graph images in PNG format. The table has the column namesname,id, andimage.
Connecting to Oracle R Enterprise in Embedded R Execution
To establish a connection to Oracle R Enterprise on the Oracle Database server during the embedded R execution, you can specify the control argument ore.connect in the parameters cursor. Doing so establishes a connection using the credentials of the user who invoked the embedded R function. It also automatically loads the ORE package. Establishing an Oracle R Enterprise connection is required to save objects in an Oracle R Enterprise R object datastore or to load objects from a datastore. It also allows you to explicitly use the Oracle R Enterprise transparency layer.
See Also:
"Optional and Control Arguments" for information on other control argumentsManaging Scripts Using the SQL API
The functions in the SQL API for embedded R execution require as an argument a named R script that is stored in the R script repository of the Oracle database. The sys.rq_scripts view contains the names and contents of the available scripts.
In the SQL API, the functions sys.rqScriptCreate and sys.rqScriptDrop create and drop scripts. As described in "Security Considerations for Scripts", to create a script or drop one from the R script repository requires the RQADMIN role.
When using the sys.rqScriptCreate function, you must specify a name for the script and an R script that contains a single R function definition. Calls to the functions sys.rqScriptCreate and sys.rqScriptDrop must be wrapped in a BEGIN-END PL/SQL block. The database stores the function as a character large object (a CLOB), so you must enclose the function definition in single quotes to specify it as a string.
The syntax of the functions is the following:
sys.rqScriptCreate (
V_NAME IN VARCHAR2
V_SCRIPT IN CLOB)
sys.rqScriptDrop (
V_NAME IN VARCHAR2)
Table 6-3 describes the parameters of the script creation and deletion functions.
Table 6-3 Parameters of sys.rqScriptCreate and sys.rqScriptDrop
| Parameter | Description |
|---|---|
|
|
The name of the script to create or drop. |
|
|
An R function definition. |
Example 6-19 first executes the sys.rqScriptDrop function to ensure that the R script repository does not contain a script with the specified name. The example then executes the sys.rqScriptCreate function to create the user-defined function named myRandomRedDots2. The user-defined function accepts two arguments, and it returns a data.frame object that has two columns and that plots the specified number of random normal values. The sys.rqScriptCreate function stores the user-defined function in the R script repository.
Example 6-19 Dropping and Creating an R Script with the SQL APIs
BEGIN
sys.rqScriptDrop('myRandomRedDots2');
END;
/
BEGIN
sys.rqScriptCreate('myRandomRedDots2',
'function(divisor = 100, numDots = 100) {
id <- 1:10
plot(1:numDots, rnorm(numDots), pch = 21, bg = "red", cex = 2 )
data.frame(id = id, val = id / divisor)
}');
END;
/
See Also:
-
"rqEval Function" for examples that use the
myRandomRedDots2script.
Managing Datastores in SQL
Oracle R Enterprise provides basic management for datastores in SQL. Basic datastore management includes show, search, and drop. The following functions and views are provided:
-
rquser_DataStoreListis a view that contains datastore-level information for all datastores in the current user schema. The information consists of datastore name, number of objects, size, creation date, and description.The following examples illustrate using the view:
SELECT * from rquser_DataStoreList; SELECT dsname, nobj, dssize FROM rquser_datastorelist WHERE dsname = 'ds_1';
-
rquser_DataStoreContentsis a view that contains object-level information about all of the datastores in the current user schema. The information consists of object name, size, class, length, number of rows and columns.The following example lists the datastore contents for datastore
datastore_1:SELECT * FROM rquser_DataStoreContents WHERE dsname = 'datastore_1';
-
rqDropDataStoredeletes a datastore and all of the objects in the datastore.Syntax:
rqDropDataStore('<ds_name>'), where<ds_name>is the name of the datastore to delete.The following example deletes the datastore
datastore_1from the current user schema:rqDropDataStore('datastore_1')
rqEval Function
The rqEval function executes the R function in the script specified by the EXP_NAM parameter. You can pass arguments to the R function with the PAR_CUR parameter.
The rqEval function does not automatically receive any data from the database. The R function generates the data that it uses or it explicitly retrieves it from a data source such as Oracle Database, other databases, or flat files.
The R function returns an R data.frame object, which appears as a SQL table in the database. You define the form of the returned value with the OUT_QRY parameter.
Table 6-4 describes the parameters of the rqEval function.
Table 6-4 Parameters of the rqEval Function
| Parameter | Description |
|---|---|
|
|
A cursor that contains argument values to pass to the R function specified by the |
|
|
One of the following:
|
|
|
The name of a script in the R script repository. |
Function rqEval returns a table that has the structure specified by the OUT_QRY parameter value.
Example 6-20 invokes the function myRandomRedDots2 that was created and stored in the R script repository in Example 6-19. The value of the first parameter to rqEval is NULL, which specifies that no arguments are supplied to the function myRandomRedDots2. The value of second parameter is a string that specifies a SQL statement that describes the column names and data types of the data.frame returned by rqEval. The value of third parameter is the name of the script in the R script repository.
SELECT * FROM table(rqEval(NULL, 'SELECT 1 id, 1 val FROM dual', 'myRandomRedDots2'));
In Oracle SQL Developer, the results of the SELECT statement are:
ID VAL
---------- ----------
1 .01
2 .02
3 .03
4 .04
5 .05
6 .06
7 .07
8 .08
9 .09
10 .1
10 rows selected
Example 6-21 provides arguments to the R function by specifying a cursor as the first parameter to rqEval. The cursor specifies multiple arguments in a single row of scalar values.
Example 6-21 Passing Arguments to the R Function invoked by rqEval
SELECT *
FROM table(rqEval(cursor(SELECT 50 "divisor", 500 "numDots" FROM dual),
'SELECT 1 id, 1 val FROM dual',
'myRandomRedDots2'));
In Oracle SQL Developer, the results of the SELECT statement are:
ID VAL
---------- ----------
1 .02
2 .04
3 .06
4 .08
5 .1
6 .12
7 .14
8 .16
9 .18
10 .2
10 rows selected
Example 6-22 creates a script named PNG_Example and stores it in the R script repository. The invocation of rqEval specifies an OUT_QRY value of 'PNG'.
Example 6-22 Specifying PNG as the Output Table Definition
BEGIN
sys.rqScriptDrop('PNG_Example');
sys.rqScriptCreate('PNG_Example',
'function(){
dat <- data.frame(y = log(1:100), x = 1:100)
plot(lm(y ~ x, dat))
}');
END;
/
SELECT *
FROM table(rqEval(NULL,'PNG','PNG_Example'));
In Oracle SQL Developer, the results of the SELECT statement are:
NAME ID IMAGE
------ ---- ------
1 (BLOB)
2 (BLOB)
3 (BLOB)
4 (BLOB)
rqGroupEval Function
The rqGroupEval function is a user-defined function that identifies a grouping column. The user defines an rqGroupEval function in PL/SQL using the SQL object rqGroupEvalImpl, which is a generic implementation of the group apply functionality in SQL. The implementation supports data-parallel execution, in which one or more R engines perform the same R function, or task, on different partitions of data. The data is partitioned according to the values of the grouping column.
Only one grouping column is supported. If you have multiple columns, then combine the columns into one column and use the new column as the grouping column.
The rqGroupEval function executes the R function in the script specified by the EXP_NAM parameter. You pass data to the R function with the INP_CUR parameter. You can pass arguments to the R function with the PAR_CUR parameter.
The R function returns an R data.frame object, which appears as a SQL table in the database. You define the form of the returned value with the OUT_QRY parameter.
To create an rqGroupEval function, you create the following two PL/SQL objects:
-
A PL/SQL package that specifies the types of the result to return.
-
A function that takes the return value of the package and uses the return value with
PIPELINED_PARALLEL_ENABLEset to indicate the column on which to partition data.
rqGroupEval (
INP_CUR REF CURSOR IN
PAR_CUR REF CURSOR IN
OUT_QRY VARCHAR2 IN
GRP_COL VARCHAR2 IN
EXP_NAM VARCHAR2 IN)
Table 6-7 describes the parameters of the user-defined rqGroupEval function.
Table 6-5 Parameters of the rqGroupEval Function
| Parameter | Description |
|---|---|
|
|
A cursor that specifies the data to pass to the R function specified by the |
|
|
A cursor that contains argument values to pass to the R function. |
|
|
One of the following:
|
|
|
The name of the grouping column by which to partition the data. |
|
|
The name of a script in the R script repository. |
The user-defined rqGroupEval function returns a table that has the structure specified by the OUT_QRY parameter value.
Example 6-23 has a PL/SQL block that drops the script myC5.0Function to ensure that the script does not exist in the R script repository. It then creates a function and stores it as the script myC5.0Function in the repository.
The R function accepts two arguments: the data on which to operate and a prefix to use in creating datastores. The function uses the C50 package to build C5.0 models on the churn data set from C50. The function builds one churn model on the data for each state.
The myC5.0Function function loads the C50 package so that the function body has access to it when the function executes in an R engine on the database server. The function then creates a datastore name using the datastore prefix and the name of a state. To exclude the state name from the model, the function deletes the column from the data.frame. Because factors in the data.frame are converted to character vectors when they are loaded in the user-defined embedded R function, the myC5.0Function function explicitly converts the character vectors back to R factors.
The myC5.0Function function gets the data for the state from the specified columns and then creates a model for the state and saves the model in a datastore. The R function returns TRUE to have a simple value that can appear as the result of the function execution.
Example 6-23 next creates a PL/SQL package, churnPkg, and a user-defined function, churnGroupEval. In defining an rqGroupEval function implementation, the PARALLEL_ENABLE clause is optional but the CLUSTER BY clause is required.
Finally, the example executes a SELECT statement that invokes the churnGroupEval function. In the INP_CUR argument of the churnGroupEval function, the SELECT statement specifies the PARALLEL hint to use parallel execution of the R function and the data set to pass to the R function. The INP_CUR argument of the churnGroupEval function specifies connecting to Oracle R Enterprise and the datastore prefix to pass to the R function. The OUT_QRY argument specifies returning the value in XML format, the GRP_NAM argument specifies using the state column of the data set as the grouping column, and the EXP_NAM argument specifies the myC5.0Function script in the R script repository as the R function to invoke.
Example 6-23 Using an rqGroupEval Function
BEGIN
sys.rqScriptDrop('myC5.0Function');
sys.rqScriptCreate('myC5.0Function',
'function(dat, datastorePrefix) {
library(C50)
datastoreName <- paste(datastorePrefix, dat[1, "state"], sep = "_")
dat$state <- NULL
dat$churn <- as.factor(dat$churn)
dat$area_code <- as.factor(dat$area_code)
dat$international_plan <- as.factor(dat$international_plan)
dat$voice_mail_plan <- as.factor(dat$voice_mail_plan)
mod <- C5.0(churn ~ ., data = dat, rules = TRUE)
ore.save(mod, name = datastoreName)
TRUE
}');
END;
/
CREATE OR REPLACE PACKAGE churnPkg AS
TYPE cur IS REF CURSOR RETURN CHURN_TRAIN%ROWTYPE;
END churnPkg;
/
CREATE OR REPLACE FUNCTION churnGroupEval(
inp_cur churnPkg.cur,
par_cur SYS_REFCURSOR,
out_qry VARCHAR2,
grp_col VARCHAR2,
exp_txt CLOB)
RETURN SYS.AnyDataSet
PIPELINED PARALLEL_ENABLE (PARTITION inp_cur BY HASH ("state"))
CLUSTER inp_cur BY ("state")
USING rqGroupEvalImpl;
/
SELECT *
FROM table(churnGroupEval(
cursor(SELECT * /*+ parallel(t,4) */ FROM CHURN_TRAIN t),
cursor(SELECT 1 AS "ore.connect",
'myC5.0model' AS "datastorePrefix" FROM dual),
'XML', 'state', 'myC5.0Function'));
For each of 50 states plus Washington, D.C., the SELECT statement returns from the churnGroupEval table function the name of the state and an XML string that contains the value TRUE.
rqRowEval Function
The rqRowEval function executes the R function in the script specified by the EXP_NAM parameter. You pass data to the R function with the INP_CUR parameter. You can pass arguments to the R function with the PAR_CUR parameter. The ROW_NUM parameter specifies the number of rows that should be passed to each invocation of the R function. The last chunk may have fewer rows than the number specified.
The rqRowEval function supports data-parallel execution, in which one or more R engines perform the same R function, or task, on disjoint chunks of data. Oracle Database handles the management and control of the potentially multiple R engines that run on the database server machine, automatically chunking and passing data to the R engines executing in parallel. Oracle Database ensures that R function executions for all chunks of rows complete, or the rqRowEval function returns an error.
The R function returns an R data.frame object, which appears as a SQL table in the database. You define the form of the returned value with the OUT_QRY parameter.
rqRowEval (
INP_CUR REF CURSOR IN
PAR_CUR REF CURSOR IN
OUT_QRY VARCHAR2 IN
ROW_NUM NUMBER IN
EXP_NAM VARCHAR2 IN)
Table 6-6 describes the parameters of the rqRowEval function.
Table 6-6 Parameters of the rqRowEval Function
| Parameter | Description |
|---|---|
|
|
A cursor that specifies the data to pass to the R function specified by the |
|
|
A cursor that contains argument values to pass to the R function. |
|
|
One of the following:
|
|
|
The number of rows to include in each invocation of the R function. |
|
|
The name of a script in the R script repository. |
Function rqRowEval returns a table that has the structure specified by the OUT_QRY parameter value.
Example 6-24 uses the C50 package to score churn data (that is, to predict which customers are likely to churn) using C5.0 decision tree models. The example scores the customers from the specified state in parallel. This example produces the same result as the invocation of function ore.rowApply in Example 6-15.
Tip:
Example 6-24 uses the CHURN_TEST table and themyXLevels datastore created by Example 6-15 so in R you should invoke the functions that create the table and that get the xlevels object and save it in the myXLevels datastore in Example 6-15 before running Example 6-24.As Example 6-23 does, Example 6-24 creates a user-defined function and saves the function in the R script repository. The user-defined function creates a C5.0 model for a state and saves the model in a datastore. In Example 6-24, however, the user-defined function myC5.0FunctionForLevels uses the list of levels created in Example 6-15 instead of computing the levels using the as.factor function as function myC5.0Function does in Example 6-23. The function myC5.0FunctionForLevels returns the value TRUE.
As Example 6-23 does, Example 6-24 creates the PL/SQL package churnPkg and the function churnGroupEval. Example 6-23 declares a cursor to get the names of the datastores that include the string myC5.0modelFL and then executes a PL/SQL block that deletes those datastores. The example next executes a SELECT statement that invokes the churnGroupEval function. The churnGroupEval function invokes the myC5.0FunctionForLevels function to generate the C5.0 models and save them in datastores.
Example 6-24 then creates the myScoringFunction function and stores it in the R script repository. The function scores a C5.0 model for the levels of a state and returns the results in a data.frame.
Finally, Example 6-24 executes a SELECT statement that invokes the rqRowEval function. The input cursor to the rqRowEval function uses the PARALLEL hint to specify the degree of parallelism to use. The cursor specifies the CHURN_TEST table as the data source and filters the rows to include only those for Massachusetts. All rows processed use the same predictive model.
The parameters cursor specifies the ore.connect control argument to connect to Oracle R Enterprise on the database server and specifies values for the datastorePrefix and xlevelsDatastore arguments to the myScoringFunction function.
The SELECT statement for the OUT_QRY parameter specifies the format of the output. The ROW_NUM parameter specifies 200 as the number of rows to process at a time in each parallel R engine. The EXP_NAME parameter specifies myScoringFunction in the R script repository as the R function to invoke.
Example 6-24 Using an rqRowEval Function
BEGIN
sys.rqScriptDrop('myC5.0FunctionForLevels');
sys.rqScriptCreate('myC5.0FunctionForLevels',
'function(dat, xlevelsDatastore, datastorePrefix) {
library(C50)
state <- dat[1,"state"]
datastoreName <- paste(datastorePrefix, dat[1, "state"], sep = "_")
dat$state <- NULL
ore.load(name = xlevelsDatastore) # To get the xlevels object.
for (j in names(xlevels))
dat[[j]] <- factor(dat[[j]], levels = xlevels[[j]])
c5mod <- C5.0(churn ~ ., data = dat, rules = TRUE)
ore.save(c5mod, name = datastoreName)
TRUE
}');
END;
/
CREATE OR REPLACE PACKAGE churnPkg AS
TYPE cur IS REF CURSOR RETURN CHURN_TEST%ROWTYPE;
END churnPkg;
/
CREATE OR REPLACE FUNCTION churnGroupEval(
inp_cur churnPkg.cur,
par_cur SYS_REFCURSOR,
out_qry VARCHAR2,
grp_col VARCHAR2,
exp_txt CLOB)
RETURN SYS.AnyDataSet
PIPELINED PARALLEL_ENABLE (PARTITION inp_cur BY HASH ("state"))
CLUSTER inp_cur BY ("state")
USING rqGroupEvalImpl;
/
DECLARE
CURSOR c1
IS
SELECT dsname FROM rquser_DataStoreList WHERE dsname like 'myC5.0modelFL%';
BEGIN
FOR dsname_st IN c1
LOOP
rqDropDataStore(dsname_st.dsname);
END LOOP;
END;
SELECT *
FROM table(churnGroupEval(
cursor(SELECT * /*+ parallel(t,4) */ FROM CHURN_TEST t),
cursor(SELECT 1 AS "ore.connect",
'myXLevels' as "xlevelsDatastore",
'myC5.0modelFL' AS "datastorePrefix" FROM dual),
'XML', 'state', 'myC5.0FunctionForLevels'));
BEGIN
sys.rqScriptDrop('myScoringFunction');
sys.rqScriptCreate('myScoringFunction',
'function(dat, xlevelsDatastore, datastorePrefix) {
library(C50)
state <- dat[1, "state"]
datastoreName <- paste(datastorePrefix, state, sep = "_")
dat$state <- NULL
ore.load(name = xlevelsDatastore) # To get the xlevels object.
for (j in names(xlevels))
dat[[j]] <- factor(dat[[j]], levels = xlevels[[j]])
ore.load(name = datastoreName)
res <- data.frame(pred = predict(c5mod, dat, type = "class"),
actual= dat$churn,
state = state)
res
}');
END;
/
SELECT * FROM table(rqRowEval(
cursor(select /*+ parallel(t, 4) */ *
FROM CHURN_TEST t
WHERE "state" = 'MA'),
cursor(SELECT 1 as "ore.connect",
'myC5.0modelFL' as "datastorePrefix",
'myXLevels' as "xlevelsDatastore"
FROM dual),
'SELECT ''aaa'' "pred",''aaa'' "actual" , ''aa'' "state" FROM dual',
200, 'myScoringFunction'));
In Oracle SQL Developer, the results of the last SELECT statement are:
pred actual state ---- ------ ----- no no MA no no MA no no MA no no MA no no MA no no MA no no MA no yes MA yes yes MA yes yes MA no no MA no no MA no no MA no no MA no no MA no no MA yes yes MA no no MA no no MA no no MA no no MA no no MA no no MA no no MA no no MA no no MA no no MA no no MA no no MA no no MA no yes MA no no MA no no MA no no MA yes yes MA no no MA no no MA no no MA 38 rows selected
rqTableEval Function
The rqTableEval function executes the R function in the script specified by the EXP_NAM parameter. You pass data to the R function with the INP_CUR parameter. You can pass arguments to the R function with the PAR_CUR parameter.
The R function returns an R data.frame object, which appears as a SQL table in the database. You define the form of the returned value with the OUT_QRY parameter.
rqTableEval (
INP_CUR REF CURSOR IN
PAR_CUR REF CURSOR IN
OUT_QRY VARCHAR2 IN
EXP_NAM VARCHAR2 IN)
Table 6-7 describes the parameters of the rqTableEval function.
Table 6-7 Parameters of the rqTableEval Function
| Parameter | Description |
|---|---|
|
|
A cursor that specifies the data to pass to the R function specified by the |
|
|
A cursor that contains argument values to pass to the input function. |
|
|
One of the following:
|
|
|
The name of a script in the R script repository. |
Function rqTableEval returns a table that has the structure specified by the OUT_QRY parameter value.
Example 6-25 first has a PL/SQL block that drops the script myNaiveBayesModel to ensure that the script does not exist in the R script repository. It then creates a function and stores it as the script myNaiveBayesModel in the repository.
The R function accepts two arguments: the data on which to operate and the name of a datastore. The function builds a Naive Bayes model on the iris data set. Naive Bayes is found in the e1071 package.
The myNaiveBayesModel function loads the e1071 package so that the function body has access to it when the function executes in an R engine on the database server. Because factors in the data.frame are converted to character vectors when they are loaded in the user-defined embedded R function, the myNaiveBayesModel function explicitly converts the character vector to an R factor.
The myNaiveBayesModel function gets the data from the specified column and then creates a model and saves it in a datastore. The R function returns TRUE to have a simple value that can appear as the result of the function execution.
Example 6-25 next executes a SELECT statement that invokes the rqTableEval function. In the INP_CUR argument of the rqTableEval function, the SELECT statement specifies the data set to pass to the R function. The data is from the IRIS table that was created by invoking ore.create(iris, "IRIS"), which is not shown in the example. The INP_CUR argument of the rqTableEval function specifies the name of a datastore to pass to the R function and specifies the ore.connect control argument to establish an Oracle R Enterprise connection to the database during the embedded R execution of the user-defined R function. The OUT_QRY argument specifies returning the value in XML format, and the EXP_NAM argument specifies the myNaiveBayesModel script in the R script repository as the R function to invoke.
Example 6-25 Using the rqTableEval Function
BEGIN
sys.rqScriptDrop('myNaiveBayesModel');
sys.rqScriptCreate('myNaiveBayesModel',
'function(dat, datastoreName) {
library(e1071)
dat$Species <- as.factor(dat$Species)
nbmod <- naiveBayes(Species ~ ., dat)
ore.save(nbmod, name = datastoreName)
TRUE
}');
END;
/
SELECT *
FROM table(rqTableEval(
cursor(SELECT * FROM IRIS),
cursor(SELECT 'myNaiveBayesDatastore' "datastoreName",
1 as "ore.connect" FROM dual),
'XML', 'myNaiveBayesModel'));
The SELECT statement returns from the rqTableEval table function an XML string that contains the value TRUE.
The myNaiveBayesDatastore datastore now exists and contains the object nbmod, as shown by the following SELECT statement.
SQL> SELECT * from rquser_DataStoreContents 2 WHERE dsname = 'myNaiveBayesDatastore'; DSNAME OBJNAME CLASS OBJSIZE LENGTH NROW NCOL --------------------- ------- ---------- ------- ------ ---- ---- myNaiveBayesDatastore nbmod naiveBayes 1485 4
In a local R session, you could load the model and display it, as in the following:
R> ore.load("myNaiveBayesDatastore")
[1] "nbmod"
R> nbmod
$apriori
Y
setosa versicolor virginica
50 50 50
$tables
$tables$Sepal.Length
Sepal.Length
Y [,1] [,2]
setosa 5.006 0.3524897
versicolor 5.936 0.5161711
virginica 6.588 0.6358796
$tables$Sepal.Width
Sepal.Width
Y [,1] [,2]
setosa 3.428 0.3790644
versicolor 2.770 0.3137983
virginica 2.974 0.3224966
$tables$Petal.Length
Petal.Length
Y [,1] [,2]
setosa 1.462 0.1736640
versicolor 4.260 0.4699110
virginica 5.552 0.5518947
$tables$Petal.Width
Petal.Width
Y [,1] [,2]
setosa 0.246 0.1053856
versicolor 1.326 0.1977527
virginica 2.026 0.2746501
$levels
[1] "setosa" "versicolor" "virginica"
$call
naiveBayes.default(x = X, y = Y, laplace = laplace)
attr(,"class")
[1] "naiveBayes"